Contents
- Cytoscape @version@ User Manual
- Introduction
- Quick Tour of Cytoscape
- Command Line Arguments
- Cytoscape Preferences
- Creating Networks
- Supported Network File Formats
- Node and Edge Attributes
- Navigation and Layout
- Finding and Filtering Nodes and Edges
- Editing Networks
- Nested Networks
- Rendering Engine
- Annotation
- Linkout
Cytoscape @version@ User Manual
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
Copyright (c) 2001-2015 The Cytoscape Consortium
Introduction
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
This manual describes the installation and use of Cytoscape. For a more thorough understanding of Cytoscape and its ecosystem, we highly recommend reading the Welcome Letter accessible on the http://cytoscape.org website.
Quick Tour of Cytoscape
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
Welcome Screen
When you start Cytoscape, you can access basic functions from the Welcome Screen:
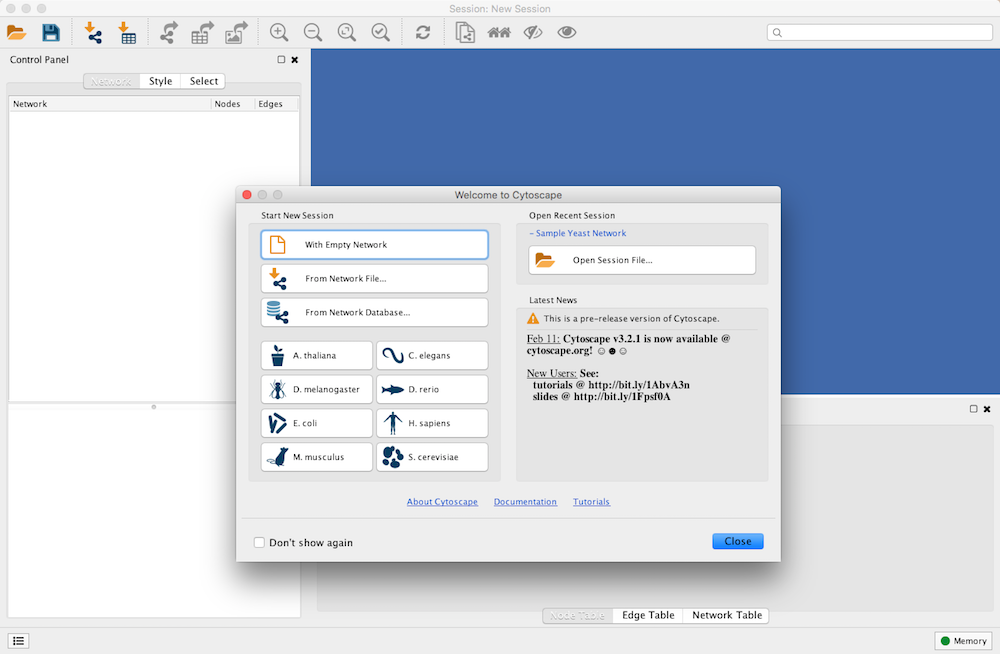
The Welcome Screen is designed to access commonly used features of Cytoscape including:
- Create new network
- Import network
- From file
- From public database
- Import interactome for model organisms
- Open recently used session file
Also, a news panel always display latest information for users. For information on user privacy, see the Cytoscape Privacy Policy.
Basic Features
When a network is loaded, Cytoscape will look similar to the image below:
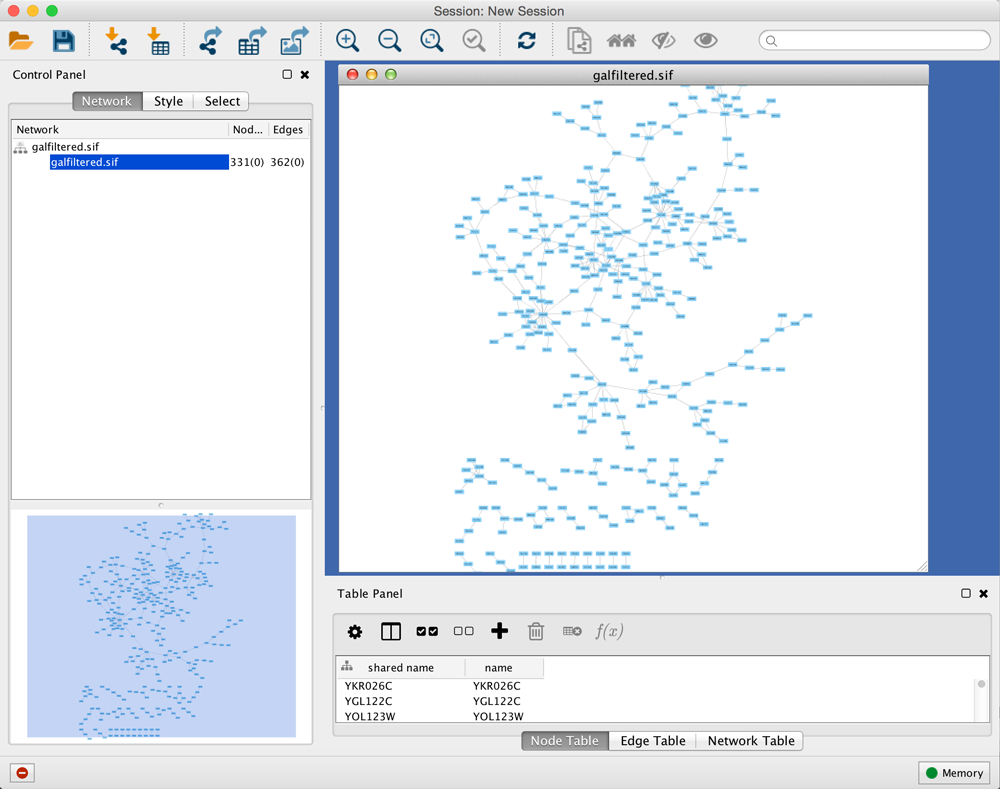
Most functionalities are self-explanatory, but we'll go through a concise explanation for clarity. The main window here has several components:
- The Menu Bar at the top (see below for more information about each menu).
- The Tool Bar, which contains icons for commonly used functions. These functions are also available via the menus. Hover the mouse pointer over an icon and wait momentarily for a description to appear as a tooltip.
- The Network Panel (Network tab of the Control Panel, top left). This contains an optional network overview pane (shown at the bottom left).
- The main Network View Window, which displays the network.
- The Table Panel (bottom right panel), which displays columns of selected nodes and edges and enables you to modify the values of column data.
The Network Panel and Table Panel are dockable tabbed Panels. You can undock any of these panels by clicking on the Float Window control  in the upper-right corner of the CytoPanel. This is useful when you want assign the network panel as much screen space as possible. To dock the window again, click the Dock Window icon
in the upper-right corner of the CytoPanel. This is useful when you want assign the network panel as much screen space as possible. To dock the window again, click the Dock Window icon  . Clicking the Hide Panel icon
. Clicking the Hide Panel icon  will hide the panel; this can be shown again by choosing View → Show and selecting the relevant panel.
will hide the panel; this can be shown again by choosing View → Show and selecting the relevant panel.
If you click this control, for example on the Table Panel, you will now have two Cytoscape windows, the main window, and a new window labeled Table Panel, similar to the one shown below. A popup will be displayed when you put the mouse pointer on a cell.
Note that Table Panel now has a Dock Window control. If you click this control, the window will dock onto the main window. For more information on the panels in Cytoscape, see the Panels section.
Network Editing
Cytoscape also has an edit functionality that enables you to build and modify networks interactively within the network canvas. To edit a network, just right-click on the open space of network window, select the menu item Add → Node, a new node will be added to the canvas. To add an edge, right click on a node, choose the menu item Edit → Add Edge. Then select the target node, a new edge will be added between the two nodes. In a similar way annotation objects can be added; pictures, shapes or textboxes; much like in MS PowerPoint or similar software. Detailed information on network editing can be found in the Editing Networks section.
The Menus
File
The File menu contains most basic file functionality: File → Open for opening a Cytoscape session file; File → New for creating a new network, either blank for editing, or from an existing network; File → Save for saving a session file; File → Import for importing data such as networks and data; and File → Export for exporting data and images. File → Export → Network View as Graphics lets you export the network in either JPEG, PDF, PNG, Post Script or SVG format.
File → Recent Session will list recently opened session files for quick access. File → Run allows you to specify a Cytoscape script file to run, and File → Print Current Network... allows printing.
Edit
The Edit menu contains Cut, Copy and Paste functions, as well as Undo and Redo functions which undo and redo edits made in the Table Panel, the Network Editor and to layout.
There are also options for creating and destroying views (graphical representations of a network) and networks (the raw network data – not yet visualized), as well as an option for deleting selected nodes and edges from the current network. All deleted nodes and edges can be restored to the network via Edit → Undo.
There are also other editing options; Remove Duplicated Edges will delete edges that are complete duplicates, keeping one edge, Remove Self-Loops removes edges that have the same source and target node, and Delete Selected Nodes and Edges... deletes a selected subset of the network. Rename Network... allows you to rename the currently selected network.
Editing preferences for properties and apps is found under Edit → Preferences → Properties.... More details on how to edit preferences can be found here.
View
The View menu allows you to display or hide the Control Panel, Table Panel, Tool Panel and the Result Panel. It also provides the control of other view-related functionality.
Select
The Select menu contains different options for selecting nodes and edges.
Layout
The Layout menu has an array of features for visually organizing the network. The features in the top portion of the network (Bundle Edges, Clear Edge Bends, Rotate, Scale, Align and Distribute) are tools for manipulating the network visualization. The bottom section of the menu lists a variety of layout algorithms which automatically lay a network out.
Apps
The Apps menu gives you access to the App Manager (Apps → App Manager) for managing (install/update/delete) your apps and may have options added by apps that have been installed. Depending on which apps are loaded, the apps that you see may be different than what appear here. The below picture shows a Cytoscape installation without installed apps.
Note: A list of available Cytoscape apps with descriptions is available online at: http://apps.cytoscape.org |
In previous versions of Cytoscape, apps were called plugins and served a similar function.
Tools
The Tools menu contains advanced features like the Command Line Dialog, Network Analyzer, Network Merge and Workflow.
Help
The Help menu allows you to launch the online help viewer and browse the table of contents for this manual (Contents).
The Citations option displays the main literature citation for Cytoscape, as well as a list of literature citations for installed apps. The list will be different depending on the set of apps you have installed.
The Help menu also allows you to connect directly to Cytoscape Help Desk and the Bug Report interface.
Network Management
Cytoscape allows multiple networks to be loaded at a time, either with or without a view. A network stores all the nodes and edges that are loaded by the user and a view displays them.
An example where a number of networks have been loaded is shown below:
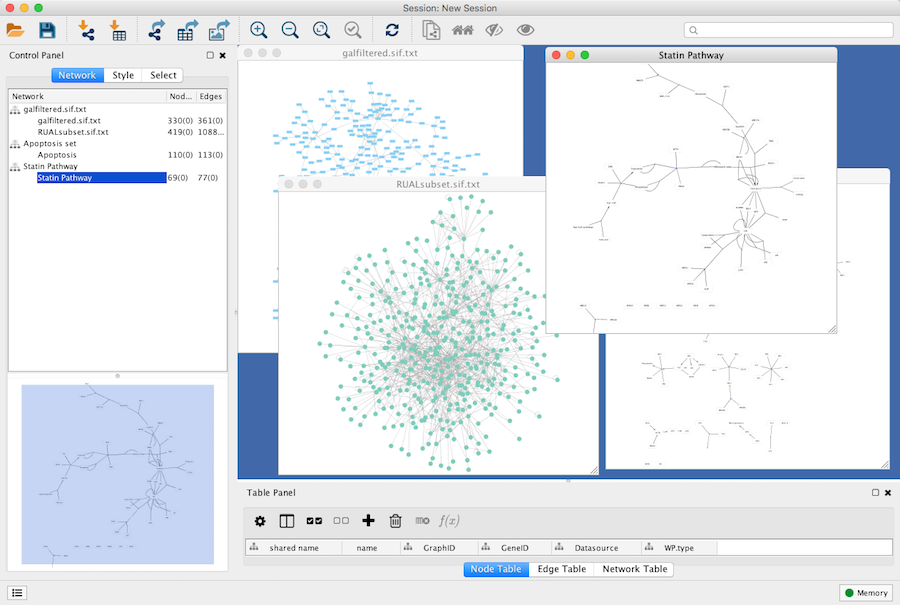
The network manager (in Control Panel) shows the networks that are loaded. Clicking on a network here will make that view active in the main window, if the view exists. Each network has a name and size (number of nodes and edges), which are shown in the network manager. If a network is loaded from a file, the network name is the name of the file.
Some networks are very large (thousands of nodes and edges) and can take a long time to display. For this reason, a network in Cytoscape may not contain a "view". Networks that have a view are in normal black font and networks that don’t have a view are highlighted in red. You can create or destroy a view for a network by right-clicking the network name in the network manager or by choosing the appropriate option in the Edit menu. You can also destroy previously loaded networks this way.
Certain operations in Cytoscape will create new networks. If a new network is created from an old network, for example by selecting a set of nodes in one network and copying these nodes to a new network (via the File → New → Network option), it will be shown immediately follows the network that it was derived from.
The available network views are also arranged as multiple overlapping windows in the network view window. You can maximize, minimize, and destroy network views by using the normal window controls for your operating system.
Arrange Network Windows
When you work on multiple networks, you can arrange the network view windows under View → Arrange Network Windows.
Vertical
Grid
The Network Overview Window
The network overview window shows an overview (or "bird’s eye view") of the network. It can be used to navigate around a large network view. The blue rectangle indicates the portion of the network currently displayed in the network view window, and it can be dragged with the mouse to view other portions of the network. Zooming in will cause the rectangle to appear smaller and vice versa.
Command Line Arguments
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
usage: cytoscape.{sh|bat} [OPTIONS]
-h,--help Print this message.
-v,--version Print the version number.
-s,--session <file> Load a cytoscape session (.cys) file.
-N,--network <file> Load a network file (any format).
-P,--props <file> Load cytoscape properties file (Java properties
format) or individual property: -P name=value.
-V,--vizmap <file> Load vizmap properties file (Cytoscape VizMap
format).
-S,--script <file> Execute commands from script file.
-R,--rest <port> Start a rest service.Any file specified for an option may be specified as either a path or as a URL. For example you can specify a network as a file (assuming that myNet.sif exists in the current working directory): cytoscape.sh -N myNet.sif.
Note: if there are spaces in the file path, be sure to put quotes around it: cytoscape.bat -N "C:\Program Files\Cytoscape\sampleData\galFiltered.sif".
Or you can specify a network as a URL: cytoscape.sh -N http://example.com/myNet.sif.
Argument |
Description |
-h,--help |
This flag generates the help output you see above and exits. |
-v,--version |
This flag prints the version number of Cytoscape and exits. |
-s,--session <file> |
This option specifies a session file to be loaded. Since only one session file can be loaded at a given time, this option may only specified once on a given command line. The option expects a .cys Cytoscape session file. It is customary, although not necessary, for session file names to contain the .cys extension. |
-N,--network <file> |
This option is used to load all types of network files. SIF, GML, and XGMML files can all be loaded using the -N option. You can specify as many networks as desired on a single command line. |
-P,--props <file> |
This option specifies Cytoscape properties. Properties can be specified either as a properties file (in Java's standard properties format), or as individual properties. To specify individual properties, you must specify the property name followed by the property value where the name and value are separated by the '=' sign. For example to specify the defaultSpeciesName: cytoscape.sh -P defaultSpeciesName=Human. If you would like to include spaces in your property, simply enclose the name and value in quotation marks: cytoscape.sh -P "defaultSpeciesName=Homo Sapiens". The property option subsumes previous options -noCanonicalization, -species, and -bioDataServer. Now it would look like: cytoscape.sh -P defaultSpeciesName=Human -P noCanonicalization=true -P bioDataServer=myServer. |
-V,--vizmap <file> |
This option specifies a Style file. |
-S,--script <file> |
This option executes commands from a specifed Cytoscape script file. |
-R,--rest <port> |
This option starts a Cytoscape REST service on the specified port. |
All options described above (except for starting a REST service) can be accessed from the menu once Cytoscape is running.
Cytoscape Preferences
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
Managing Properties
The Cytoscape properties editor, accessed via Edit → Preferences → Properties…, is used to specify default properties. Any changes made to these properties will be saved in .props files under the CytoscapeConfiguration subdirectory of the user's home directory.
Cytoscape properties are configurable using the Add, Modify and Delete buttons as seen below.
App properties may also be edited in the same way as editing Cytoscape properties. For example, to edit the properties of Linkout, select 'linkout' from the combobox of the Preferences Editor. Some apps may store properties inside session files in addition to (or instead of) storing them in the CytoscapeConfiguration directory.
Managing Bookmarks
Cytoscape contains a pre-defined list of bookmarks, which point to sample network files located on the Cytoscape web server. Users may add, modify, and delete bookmarks through the Bookmark manager, accessed by going to Edit → Preferences → Bookmarks….
There are currently several types of bookmarks (based on data categories), including network and table. Network bookmarks are URLs pointing to Cytoscape network files. These are normal networks that can be loaded into Cytoscape. Table bookmarks are URLs pointing to data table files.
Managing Proxy Servers
You can define and configure a proxy server for Cytoscape by going to Edit → Preferences → Proxy Settings….
After the proxy server is set, all network traffic related to loading a network from URL will pass through the proxy server. Cytoscape apps use this capability as well. The proxy settings are saved in cytoscape3.props. Each time you click the OK button after making a change to the proxy settings, an attempt is made to connect to a well known site on the Internet (e.g., google.com) using your settings. For both success and failure you are notified and for failure you are given an opportunity to change your proxy settings.
If you no longer need to use a proxy to connect to the Internet, simply set the Proxy type to "direct" and click the OK button.
Managing Group View
The configuration of Cytoscape group view may also be edited through Edit → Preferences → Group Preferences….
Creating Networks
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
- Importing pre-existing, fixed-format network files.
- Importing pre-existing, unformatted text or Excel files.
- Importing data from from public databases.
- Creating an empty network and manually adding nodes and edges.
Import Fixed-Format Network Files
Network files can be specified in any of the formats described in the Supported Network Formats section. Networks are imported into Cytoscape through the File → Import → Network menu. The network file can either be located directly on the local computer, or found on a remote computer (in which case it will be referenced with a URL).
Load Networks from Local Computer
In order to load a network from a local file you can select File → Import → Network → File... or click on ![]() on the tool bar. Choose the correct file in the file chooser dialog and press Open. Some sample network files of different types have been included in the sampleData folder in Cytoscape.
on the tool bar. Choose the correct file in the file chooser dialog and press Open. Some sample network files of different types have been included in the sampleData folder in Cytoscape.
After you choose a network file, another dialog will pop up. Here, you can choose either to create a new network collection for the new network, or load the new network into an existing network collection. When you choose the latter, make sure to choose the right mapping column to map the new network to the existing network collection.
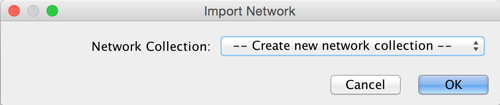
Network files in SIF, GML, and XGMML formats may also be loaded directly from the command line using the –N option.
Load Networks from a Remote Computer (URL import)
To load a network from a remote file, you can select File → Import → Network → URL.... In the import network dialog, insert the appropriate URL, either manually or using URL bookmarks. Bookmarked URLs can be accessed by clicking on the arrow to the right of the text field (see the Bookmark Manager in Preferences for more details on bookmarks). Also, you can drag and drop links from a web browser to the URL text box. Once a URL has been specified, click on the OK button to load the network.
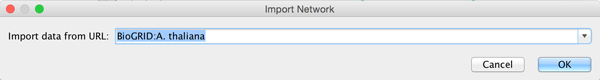
Another issue for network import is the presence of firewalls, which can affect which files are accessible to a computer. To work around this problem, Cytoscape supports the use of proxy servers. To configure a proxy server, go to Edit → Preferences→ Proxy Settings.... This is further described in the Preferences section.
Import Networks from Unformatted Table Files
Cytoscape supports the import of networks from delimited text files and Excel workbooks using File → Import → Network → File.... An interactive GUI allows users to specify parsing options for specified files. The screen provides a preview that shows how the file will be parsed given the current configuration. As the configuration changes, the preview updates automatically. In addition to specifying how the file will be parsed, the user must also choose the columns that represent the source and target nodes as well as an optional edge interaction type.
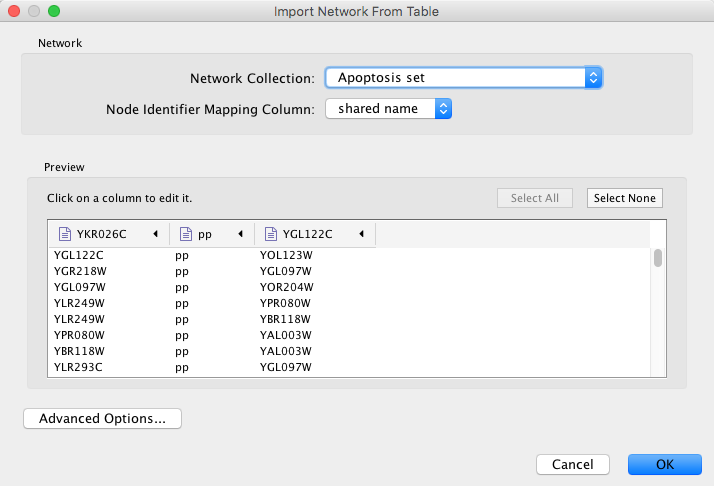
Supported Files
The Import Network from Table function supports delimited text files and Microsoft Excel Workbooks. For Excel Workbooks with multiple sheets, one sheet can be selected for import at a time. The following is a sample table file:
source target interaction boolean data string data floating point data YJR022W YNR053C pp TRUE abcd12371 1.2344543 YER116C YDL013W pp TRUE abcd12372 1.2344543 YNL307C YAL038W pp FALSE abcd12373 1.2344543 YNL216W YCR012W pd TRUE abcd12374 1.2344543 YNL216W YGR254W pd TRUE abcd12375 1.2344543
The network table files should contain at least two columns for creating network with edges. If the file has only one column, the created network will not contain any edges. The interaction type is optional in this format. Therefore, a minimal network table looks like the following:
YJR022W YNR053C YER116C YDL013W YNL307C YAL038W YNL216W YCR012W YNL216W YGR254W
One row in a network table file represents an edge and its edge data columns. This means that a network file is considered a combination of network data and edge column data. A table may contain columns that aren't meant to be edge data. In this case, you can choose not to import those columns by clicking on the column header in the preview window. This function is useful when importing a data table like the following (1):

This data file is a tab-delimited text file and contains network data (interactions), edge data, and node data. To import network and edge data from this table, choose Unique ID A as source, Unique ID B as target, and Interactor types as interaction type. Next, turn off columns used for node data (Alternative ID A, species B, etc.). Other columns can be imported as edge data.
The network import function cannot import node table columns - only edge table columns. To import node table columns from this table, please see the Node and Edge Column Data section of this manual.
Note (1): This data is taken from the A merged human interactome datasets by Andrew Garrow, Yeyejide Adeleye and Guy Warner (Unilever, Safety and Environmental Assurance Center, 12 October 2006). Actual data files are available at http://wiki.cytoscape.org/Data_Sets/
Basic Operations
To import network from text/Excel tables, please follow these steps:
Select File → Import → Network → File... or click on
 on the tool bar.
on the tool bar. - Select a table file in the file chooser dialog.
- Define the interaction parameters by specifying which columns of data contain the Source Interaction, Target Interaction, and Interaction Type. Clicking on any column header will bring up the interface for selecting source, interaction and target:
- (Optional) Define edge table columns, if applicable. Network table files can have edge table columns in addition to network data.
- Click the OK button.
Import List of Nodes Without Edges
The table import feature supports lists of nodes without edges. If you select only a source column, it creates a network without interactions. This feature is useful with the node expansion function available from some web service clients. Please read the section Importing Networks from External Database for more detail.
Advanced Options
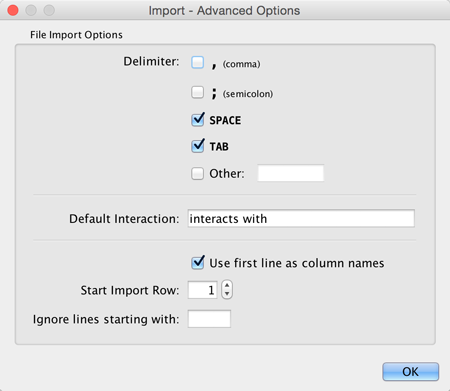
You can select several options by clicking the Advanced Options button in the main import interface.
- Delimiter: You can select multiple delimiters for text tables. By default, Tab and Space are selected as delimiters.
- Default Interaction
- Transfer first line as column names: Selecting this option will cause all edge columns to be named according to the first data entry in that column.
- Start Import Row: Set which row of the table to begin importing data from. For example, if you want to skip the first 3 rows in the file, set 4 for this option.
- Ignore lines starting with: Rows starting with this character will not be imported. This option can be used to skip comment lines in text files.
Modify Column Name/Type
In the Import Network from Table interface, you can change the name and data type of column by clicking on any column header:
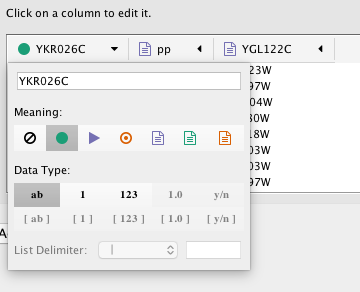
Column names and data types can be modified here.
- Modify Column Name - just enter a new column name.
- Modify Column Data Type - The following column data types are supported:
- String
- Boolean (True/False)
- Integer
- Floating Point
- List of (one of) String/Boolean/Integer/Floating Point
Cytoscape has a basic data type detection function that automatically suggests the column data type according to its entries. This can be overridden by selecting the appropriate data type from the radio buttons provided. For lists, a global delimiter must be specified (i.e., all cells in the table must use the same delimiter).
Import Networks from Public Databases
Cytoscape has a feature called Import Network from Public Databases. Users can access various kinds of databases through this function, File → Import → Network → Public Databases....
What is a Web Service?
A web service is a standardized, platform-independent mechanism for computers to interact over the internet. These days, many major biological databases publish their data with a web service API:
List of Biological Web Services: http://taverna.sourceforge.net/services
Web Services at the EBI: http://www.ebi.ac.uk/Tools/webservices/
Cytoscape core developer team have developed several web service clients using this framework. Cytoscape supports many web services including:
PSICQUIC: Standard web service for biological interaction data sets. The full list of PSICQUIC-compatible databases is available here.
The following sections describe how to import network from external databases.
Getting Started
To get started, select File → Import → Network → Public Databases....
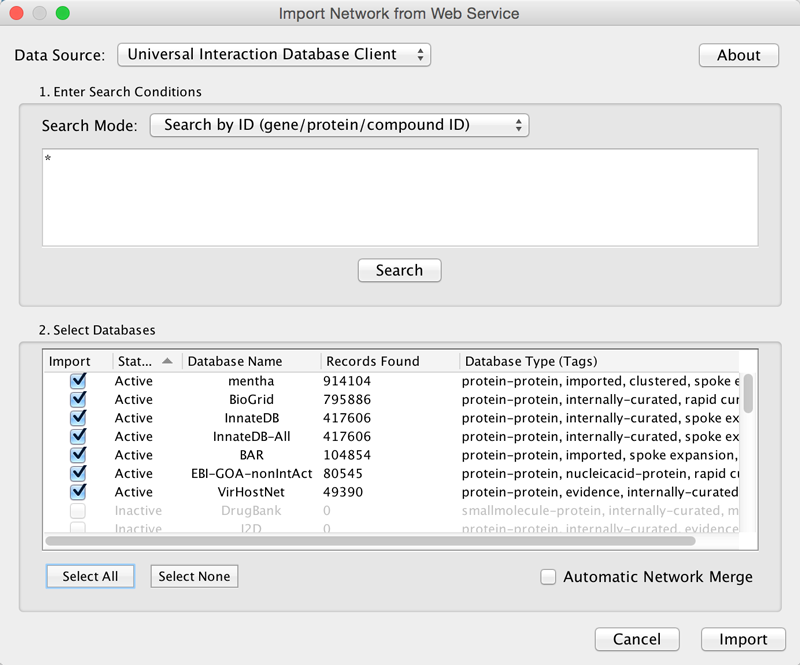
Example: Retrieving Protein-Protein Interaction Networks from Multiple Databases
Select File → Import → Network → Public Databases....
From the pull-down menu, select the Interaction databases Universal Client.
- Enter one or more search terms, such as BRCA1.
Click the Search button to start the search.
- Select databases from the hits. This selection will be saved as your default database list.
Click the Import button to import selected network data.
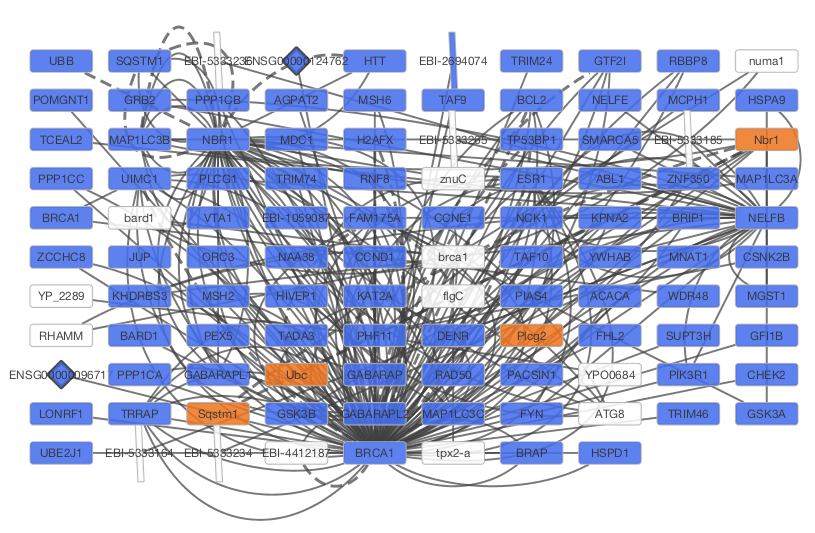
After confirming the download of interaction data, the network of BRCA1 will be imported and visualized.
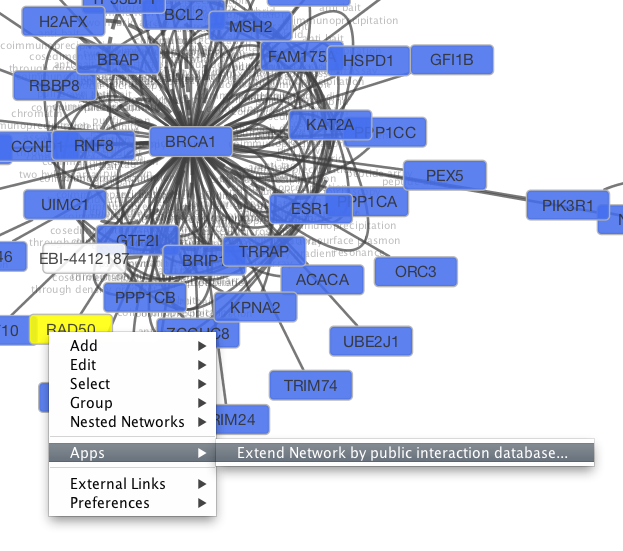
Tip: Expanding the Network: Several of the Cytoscape web services provide additional options in the node context menu. To access these options, right-click on a node and select Apps → Extend Network by public interaction database.... For example, in the screenshot, we have loaded the BRCA1 network from IntAct, and have chosen to merge this node's neighbors into the existing network.
PSICQUIC Options
PSICQUIC Web Service Client has three search modes:
- Search by ID
- Search by MIQL
- Search by Species
By default, search mode is set to Search by ID. You can search all databases by ID, such as gene symbol, Uniprot ID, or NCBI gene ID. If the search mode is set to MIQL, you can use MIQL for search. If you want to search interactions by keywords or specific functions, this is the powerful query language to filter the result. The last option is for importing all interactions for the species (i.e., interactome).
Create a New Network Manually
A new, empty network can also be created and nodes and edges manually added. To create an empty network, go to File → New → Network → Empty Network, and then manually add network components by right clicking on the network canvas or on a node.
Adding a Node
To add a new node, right-click on an empty space of the network view panel. Select Add → Node item from the pop-up menu.
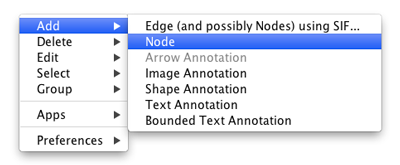
Adding an Edge
To add an edge to connect nodes, right-click on the source node. Select Edit → Add Edge from the pop-up menu. Next, click on the target node. The Images below show the two steps for drawing an edge between two nodes. You can abort the drawing of the edge by pressing Esc key. You can also select two or more nodes to connect and in the right-click menu select Add → Edges Connecting Selected Nodes to create edges connecting all selected nodes.
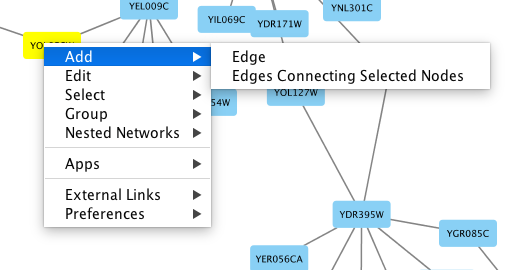
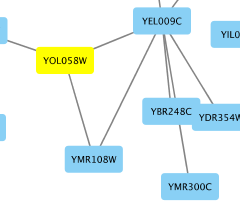
You can delete nodes and edges by selecting a number of nodes and edges, then selecting Edit → Cut. You can also delete selected nodes and edges from the Edit menu, under Edit → Delete Selected Nodes and Edges.... You can recover any nodes and edges deleted from a network by going to Edit → Undo.
Grouping nodes
Any number of nodes can be grouped together and displayed as either one group node or as the individual nodes. To create a group, select two or more nodes and right-click to select Group → Group Selected Nodes. You will be prompted to select a name for the group node. Once a group is created, you can use the right-click menu to collapse or expand the group. You can also quickly collapse/expand a group by double-clicking on the group node or any of its children to toggle back and forth.
Collapsed group
Expanded group
Adding Network Annotations
Annotations in the form of text, images or shapes can be added to the network canvas by right-clicking anywhere on the canvas and selecting one of the Annotation choices in the Add menu. You can add an image of your own, choose from a shapes library or add either plain or bounded text. Shapes and text are customizable and any added annotations can be edited from the right-click context menu.
Supported Network File Formats
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
- Simple interaction file (SIF or .sif format)
- Nested network format (NNF or .nnf format)
- Graph Markup Language (GML or .gml format)
- XGMML (extensible graph markup and modelling language).
- SBML
- BioPAX
- PSI-MI Level 1 and 2.5
- GraphML
- Delimited text
- Excel Workbook (.xls, .xlsx)
The SIF format specifies nodes and interactions only, while other formats store additional information about network layout and allow network data exchange with a variety of other network programs and data sources. Typically, SIF files are used to import interactions when building a network for the first time, since they are easy to create in a text editor or spreadsheet. Once the interactions have been loaded and network layout has been performed, the network may be saved to GML or XGMML format for interaction with other systems. All file types listed (except Excel) are text files and you can edit and view them in a regular text editor.
SIF Format
The simple interaction format is convenient for building a graph from a list of interactions. It also makes it easy to combine different interaction sets into a larger network, or add new interactions to an existing data set. The main disadvantage is that this format does not include any layout information, forcing Cytoscape to re-compute a new layout of the network each time it is loaded.
Lines in the SIF file specify a source node, a relationship type (or edge type), and one or more target nodes:
nodeA <relationship type> nodeB nodeC <relationship type> nodeA nodeD <relationship type> nodeE nodeF nodeB nodeG ... nodeY <relationship type> nodeZ
A more specific example is:
node1 typeA node2 node2 typeB node3 node4 node5 node0
The first line identifies two nodes, called node1 and node2, and a single relationship between node1 and node2 of type typeA. The second line specifies three new nodes, node3, node4, and node5; here "node2" refers to the same node as in the first line. The second line also specifies three relationships, all of type typeB and with node2 as the source, with node3, node4, and node5 as the targets. This second form is simply shorthand for specifying multiple relationships of the same type with the same source node. The third line indicates how to specify a node that has no relationships with other nodes. This form is not needed for nodes that do have relationships, since the specification of the relationship implicitly identifies the nodes as well.
Duplicate entries are ignored. Multiple edges between the same nodes must have different edge types. For example, the following specifies two edges between the same pair of nodes, one of type xx and one of type yy:
node1 xx node2 node1 xx node2 node1 yy node2
Edges connecting a node to itself (self-edges) are also allowed:
node1 xx node1
Every node and edge in Cytoscape has a name column. For a network defined in SIF format, node names should be unique, as identically named nodes will be treated as identical nodes. The name of each node will be the name in this file by default (unless another string is mapped to display on the node using styles). This is discussed in the section on Styles. The name of each edge will be formed from the name of the source and target nodes plus the interaction type: for example, sourceName (edgeType) targetName.
The tag <edgeType> can be any string. Whole words or concatenated words may be used to define types of relationships, e.g. geneFusion, cogInference, pullsDown, activates, degrades, inactivates, inhibits, phosphorylates, upRegulates, etc.
Some common interaction types used in the Systems Biology community are as follows:
pp .................. protein – protein interaction pd .................. protein -> DNA (e.g. transcription factor binding upstream of a regulating gene.)
Some less common interaction types used are:
pr .................. protein -> reaction rc .................. reaction -> compound cr .................. compound -> reaction gl .................. genetic lethal relationship pm .................. protein-metabolite interaction mp .................. metabolite-protein interaction
Delimiters
Whitespace (space or tab) is used to delimit the names in the simple interaction file format. However, in some cases spaces are desired in a node name or edge type. The standard is that, if the file contains any tab characters, then tabs are used to delimit the fields and spaces are considered part of the name. If the file contains no tabs, then any spaces are delimiters that separate names (and names cannot contain spaces).
If your network unexpectedly contains no edges and node names that look like edge names, it probably means your file contains a stray tab that's fooling the parser. On the other hand, if your network has nodes whose names are half of a full name, then you probably meant to use tabs to separate node names with spaces.
Networks in simple interactions format are often stored in files with a .sif extension, and Cytoscape recognizes this extension when browsing a directory for files of this type.
NNF
The NNF format is a very simple format that unlike SIF allows the optional assignment of single nested network per node. No other node columns can be specified. There are only 2 possible line formats:
- A node "node" contained in a "network:"
network node
- 2 nodes linked together contained in a network:
network node1 interaction node2
If a network name (first entry on a line) appeared previously as a node name (in columns 2 or 4), the network will be nested in the node with the same name. Also, if a name that has been previously defined as a network (by being listed in the first column), later appears as a node name (in columns 2 or 4), the previously defined network will be nested in the node with the same name. In summary: any time a name is used as both, a network name , and a node name, this implies that the network will be nested in the node of the same name. Additionally comments may be included on all lines. Comments start with a hash mark '#' and continue to the end of a line. Trailing comments (after data lines) and entirely blank lines anywhere are also permissible. Please note that if you load multiple NNF files in Cytoscape they will be treated like a single, long concatenated NNF file! If you need to embed spaces, tabs or backslashes in a name, you must escape it by preceding it with a backslash, so that, e.g. an embedded backslash becomes two backslashes, an embedded space a backslash followed by a space etc.
Examples
Example 1
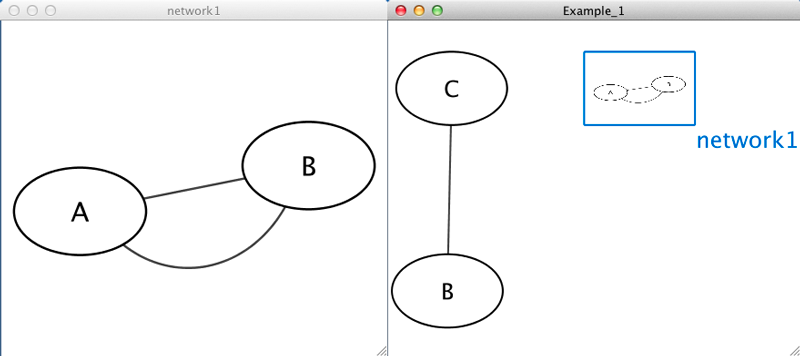
Example_1 C Example_1 network1 network1 A pp B network1 B pp A Example_1 C pp B
Example 2
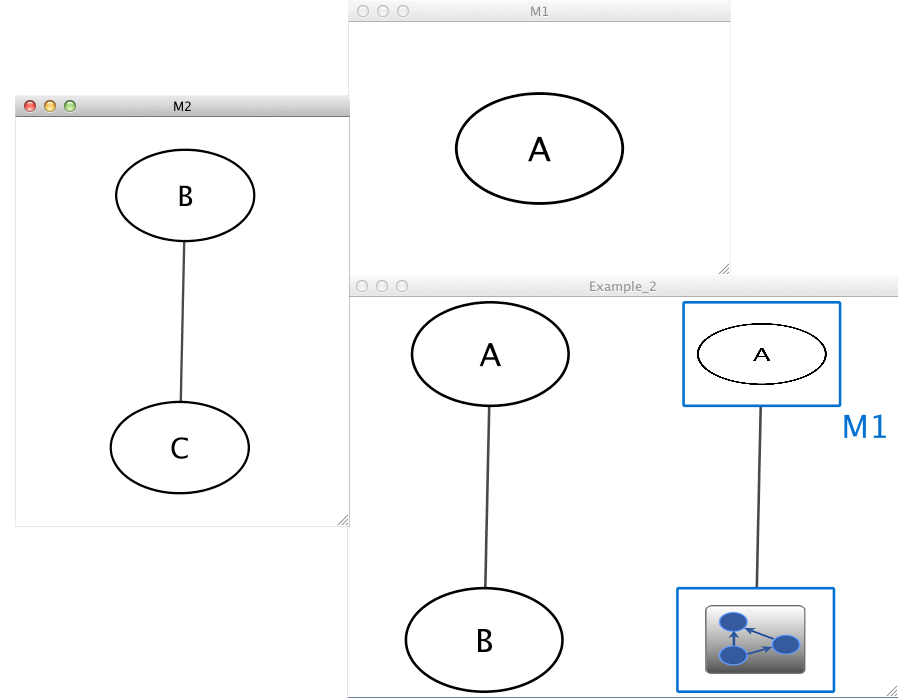
Example_2 M1 Example_2 M2 M1 A M2 B pp C Example_2 A pp B Example_2 M1 im M2
Example 3
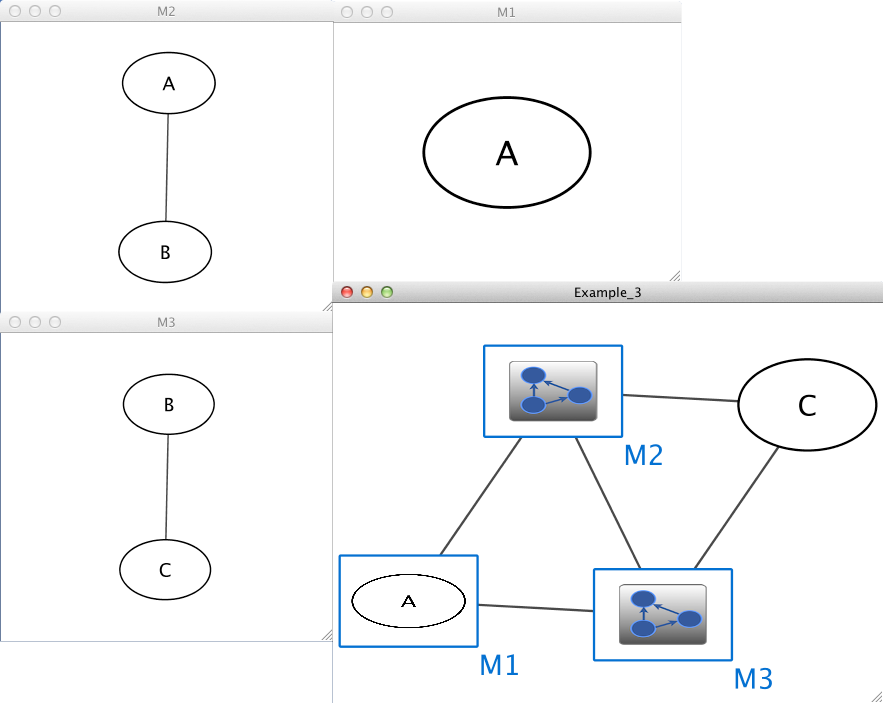
Example_3 M1 im M2 Example_3 M3 im M1 Example_3 M2 im M3 Example_3 C pp M3 Example_3 M2 pp C M1 A M2 A pp B M3 B pp C
==== Example 4 ==== 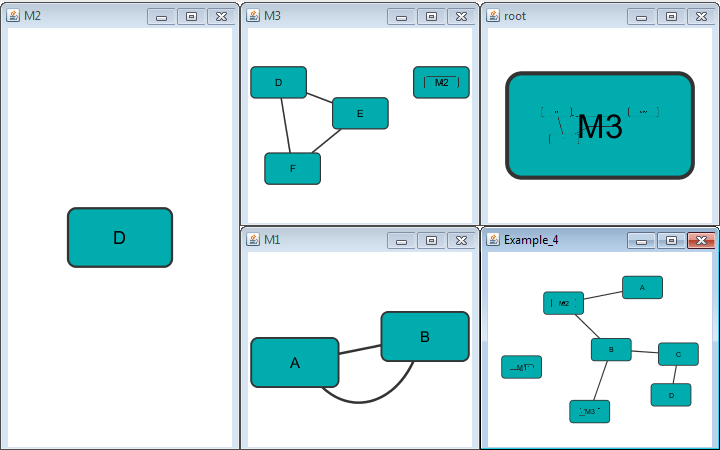 Example_4 M1*/ /* root M3*/ /* M1 A pp B M1 B pp A Example_4 C pp B M3 M2 M2 D M3 E pp F M3 D pp F M3 D pp E Example_4 D pp C Example_4 A pp M2*/ /* Example_4 B pp M3 Example_4 M2 pp B
Example_4 M1*/ /* root M3*/ /* M1 A pp B M1 B pp A Example_4 C pp B M3 M2 M2 D M3 E pp F M3 D pp F M3 D pp E Example_4 D pp C Example_4 A pp M2*/ /* Example_4 B pp M3 Example_4 M2 pp B
Example 5
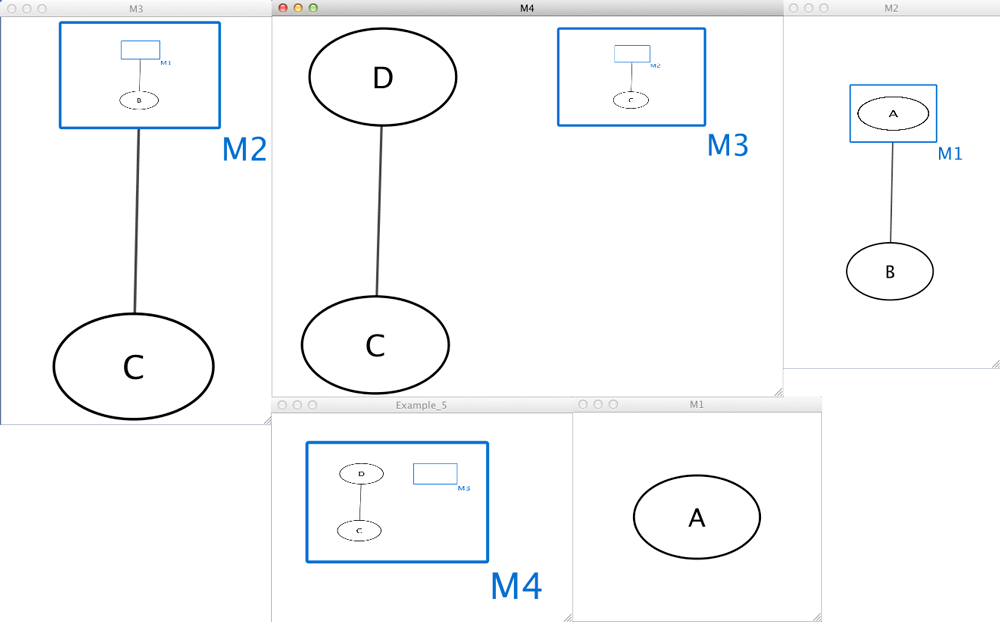
Example_5 M4 M4 D M4 M3 M3 M2 pp C M2 M1 pp B M1 A M4 C pp D
GML Format
In contrast to SIF, GML is a rich graph format language supported by many other network visualization packages. The GML file format specification is available at:
http://www.infosun.fmi.uni-passau.de/Graphlet/GML/
It is generally not necessary to modify the content of a GML file directly. Once a network is built in SIF format and then laid out, the layout is preserved by saving to and loading from GML. Properties specified in a GML file will result in a new style named Filename.style when that GML file is loaded.
XGMML Format
XGMML is the XML evolution of GML and is based on the GML definition. In addition to network data, XGMML contains node/edge/network column data. The XGMML file format specification is available at:
http://cgi5.cs.rpi.edu/research/groups/pb/punin/public_html/XGMML/
XGMML is now preferred to GML because it offers the flexibility associated with all XML document types. If you're unsure about which to use, choose XGMML.
There is a java system property "cytoscape.xgmml.repair.bare.ampersands" that can be set to "true" if you have experience trouble reading older files.
This should only be used when an XGMML file or session cannot be read due improperly encoded ampersands, as it slows down the reading process, but this is still preferable to attempting to fix such files using manual editing.
SBML (Systems Biology Markup Language) Format
The Systems Biology Markup Language (SBML) is an XML format to describe biochemical networks. SBML file format specification is available at:
BioPAX (Biological PAthways eXchange) Format
BioPAX is an OWL (Web Ontology Language) document designed to exchange biological pathways data. The complete set of documents for this format is available at:
PSI-MI Format
The PSI-MI format is a data exchange format for protein-protein interactions. It is an XML format used to describe PPI and associated data. PSI-MI XML format specification is available at:
http://psidev.sourceforge.net/mi/xml/doc/user/
GraphML
GraphML is a comprehensive and easy-to-use file format for graphs. It is based on XML. The complete set of documents for this format is available at:
http://graphml.graphdrawing.org/
Delimited Text Table and Excel Workbook
Cytoscape has native support for Microsoft Excel files (.xls, .xlsx) and delimited text files. The tables in these files can have network data and edge columns. Users can specify columns containg source nodes, target nodes, interaction types, and edge columns during file import. Some of the other network analysis tools, such as igraph (http://cneurocvs.rmki.kfki.hu/igraph/), has feature to export graph as simple text files. Cytoscape can read these text files and build networks from them. For more detail, please read the Import Free-Format Tables section of the Creating Networks section.
Cytoscape.js JSON
From Cytoscape 3.1.0 on, Cytoscape supports Cytoscape.js JSON files. You can use this feature to export your network visualizations to web browsers. Cytoscape.js has two ways to represent network data, and currently both reader and writer support only the array style graph notation. For example, this network in Cytoscape:
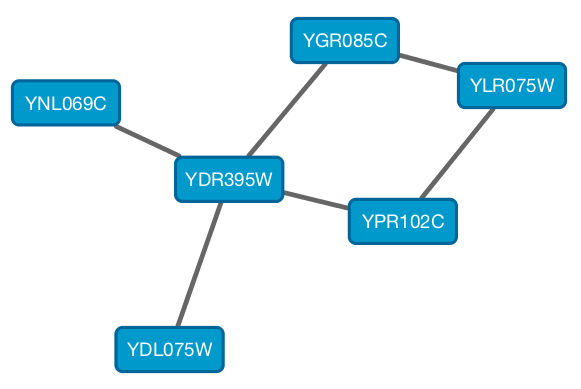
will be exported to this JSON:
{
"elements" : {
"nodes" : [ {
"data" : {
"id" : "723",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RPL31A", "RPL34", "S000002233", "ribosomal protein L31A (L34A) (YL28)" ],
"shared_name" : "YDL075W",
"SUID" : 723,
"degree_layout" : 1,
"name" : "YDL075W"
},
"position" : {
"x" : 693.0518315633137,
"y" : -49.47506554921466
},
"selected" : false
}, {
"data" : {
"id" : "726",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RP23", "RPL16B", "S000005013", "ribosomal protein L16B (L21B) (rp23) (YL15)" ],
"shared_name" : "YNL069C",
"SUID" : 726,
"degree_layout" : 1,
"name" : "YNL069C"
},
"position" : {
"x" : 627.3147710164387,
"y" : -205.99251969655353
},
"selected" : false
}, {
"data" : {
"id" : "658",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RPL11B", "S000003317", "ribosomal protein L11B (L16B) (rp39B) (YL22)" ],
"shared_name" : "YGR085C",
"SUID" : 658,
"degree_layout" : 2,
"name" : "YGR085C"
},
"position" : {
"x" : 804.3092778523762,
"y" : -245.6235926946004
},
"selected" : false
}, {
"data" : {
"id" : "660",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "KAP108", "S000002803", "SXM1" ],
"shared_name" : "YDR395W",
"SUID" : 660,
"degree_layout" : 8,
"name" : "YDR395W"
},
"position" : {
"x" : 730.8733342488606,
"y" : -157.50702317555744
},
"selected" : false
}, {
"data" : {
"id" : "579",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RPL11A", "S000006306", "ribosomal protein L11A (L16A) (rp39A) (YL22)" ],
"shared_name" : "YPR102C",
"SUID" : 579,
"degree_layout" : 2,
"name" : "YPR102C"
},
"position" : {
"x" : 841.1395696004231,
"y" : -130.77909119923908
},
"selected" : false
}, {
"data" : {
"id" : "578",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "GRC5", "QSR1", "RPL10", "S000004065", "ribosomal protein L10" ],
"shared_name" : "YLR075W",
"SUID" : 578,
"degree_layout" : 2,
"name" : "YLR075W"
},
"position" : {
"x" : 910.3755162556965,
"y" : -217.0562556584676
},
"selected" : false
} ],
"edges" : [ {
"data" : {
"id" : "659",
"source" : "658",
"target" : "578",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YGR085C (pp) YLR075W",
"SUID" : 659,
"name" : "YGR085C (pp) YLR075W"
},
"selected" : false
}, {
"data" : {
"id" : "661",
"source" : "658",
"target" : "660",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YGR085C (pp) YDR395W",
"SUID" : 661,
"name" : "YGR085C (pp) YDR395W"
},
"selected" : false
}, {
"data" : {
"id" : "724",
"source" : "660",
"target" : "723",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YDR395W (pp) YDL075W",
"SUID" : 724,
"name" : "YDR395W (pp) YDL075W"
},
"selected" : false
}, {
"data" : {
"id" : "733",
"source" : "660",
"target" : "579",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YDR395W (pp) YPR102C",
"SUID" : 733,
"name" : "YDR395W (pp) YPR102C"
},
"selected" : false
}, {
"data" : {
"id" : "727",
"source" : "660",
"target" : "726",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YDR395W (pp) YNL069C",
"SUID" : 727,
"name" : "YDR395W (pp) YNL069C"
},
"selected" : false
}, {
"data" : {
"id" : "580",
"source" : "578",
"target" : "579",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YLR075W (pp) YPR102C",
"SUID" : 580,
"name" : "YLR075W (pp) YPR102C"
},
"selected" : false
} ]
}
}And this is a sample visualization in Cytoscape.js:
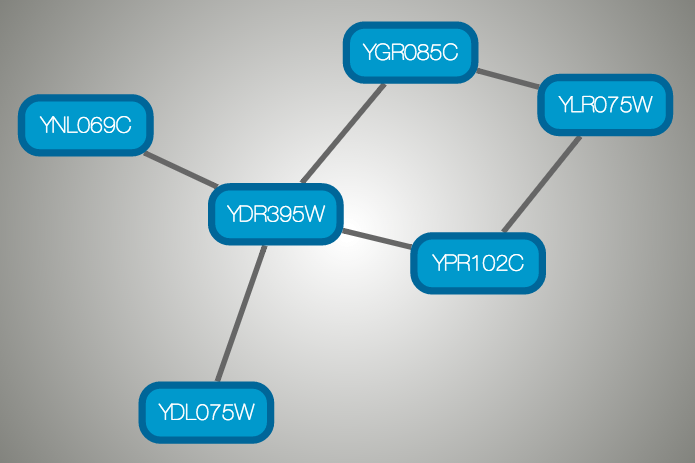
Important Note
Export network and table to Cytoscape.js feature in Cytoscape creates a JSON file WITHOUT style. This means that you need to export the style in a separate JSON file if you apply style to your network. Please read Style section for more details.
Node and Edge Attributes
Interaction networks are useful as stand-alone models. However, they are most powerful for answering scientific questions when integrated with additional information.
Cytoscape allows the user to add arbitrary node, edge and network data to Cytoscape through a data table. This could include, for example, expression data of a gene or confidence values in a protein-protein interaction. In the data table, information is linked to nodes, edges or networks by mapping the columns to one of their identifiers. Through the Table Panel the values can be further manipulated through the use of column functions and equations
A second type of data that can be associated with networks is ontology data: organized sets of controlled vocabulary terms. Because this type of data is mostly hierarchically organized, this requires a special importing facility, described in the second part of this section.
Data associated with the network elements can be visualized in a user-defined way by setting up a mapping from data in the columns to network properties (colors, shapes, and so on). The section on styles discusses this in greater detail.
Table data
Cytoscape supports several tabular formats. Of course the regular text and table formats: .tsv, .tab, .csv, .txt (comma, tab or any delimiter separated values file), .xls, .xlsx (Microsoft Excel file format). Note that for Excel file formats, only the first sheet of a workbook is currently imported. The legacy native Cytoscape formats: .attrs and .pvals (Cytoscape expression matrix) are also supported (for a more thorough discussion of these formats consult the 2.x manual). For most users the regular data table importing functionality will be sufficient; in case a format is unknown, renaming the file to a .txt extension and experimenting with delimiter and header settings in the import interface will work in most cases.
Importing data
Basic table file
The basic file format consists of a table containing at least one column with identifiers (unique names) for the nodes, edges or networks, and one or (any number) more columns with data you want to associate with these network elements.
Sample Data Table 1
Yeast Key |
Degree |
YER054C |
85 |
YBL079W |
7 |
YLR345W |
1 |
To import such a file:
Select File → Import → Table → File...( or URL... if your source data file is accessible through the internet)
- Select a data file in the file chooser panel (or enter the URL in the displayed box). This file can be in any of the accepted formats mentioned above.
In the Import Columns from Table panel, select the Importing Type. Cytoscape can import table columns to node, edge, and network table columns.
(Optional) Choose if you would like to import the file for all of the available networks or only selected networks using the check box in the expandable Network Options" panel (this panel is collapsed by default). Select networks from the list.
- (Optional) If the table is not properly delimited in the preview panel, change the delimiter in the Text File Import Options panel (the default delimiter is Tab).
- By default, the first column is designated as the primary key. Change the key column if necessary, see below for an example.
- The data attribute column name can be changed by right clicking on the header. A dialog appears that enables specification of the datatype and column name.
- Left clicking selects or deselects the column for importing.
- Click the Import button.
The user interface of the "Import Column from Table" window is similar to that of the "Import Network from Table" window.
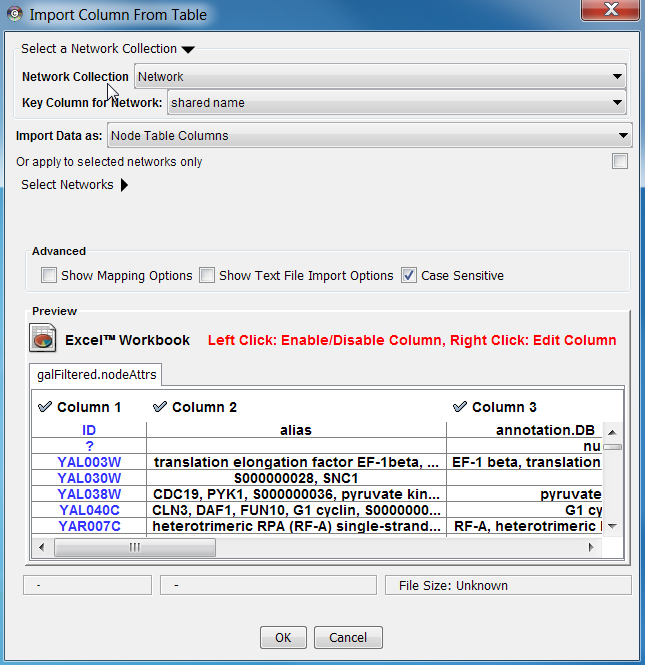
If the data only relates to a specific network select the 'Apply to selected networks only' box and select the specific network from the list (click 'Select Networks' to show this list)
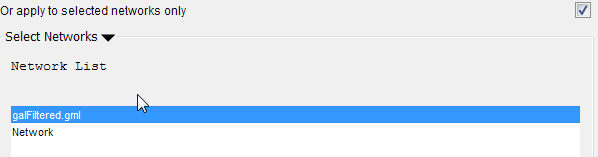
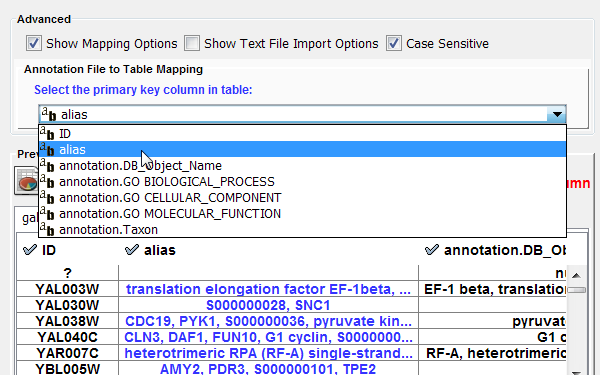
Mapping data options
In Cytoscape 3.0 data columns with primitive data types (string, boolean, floating point, and integer) can be selected as the key column using the dropdown list provided. Complex data attributes such as lists are not supported as keys.
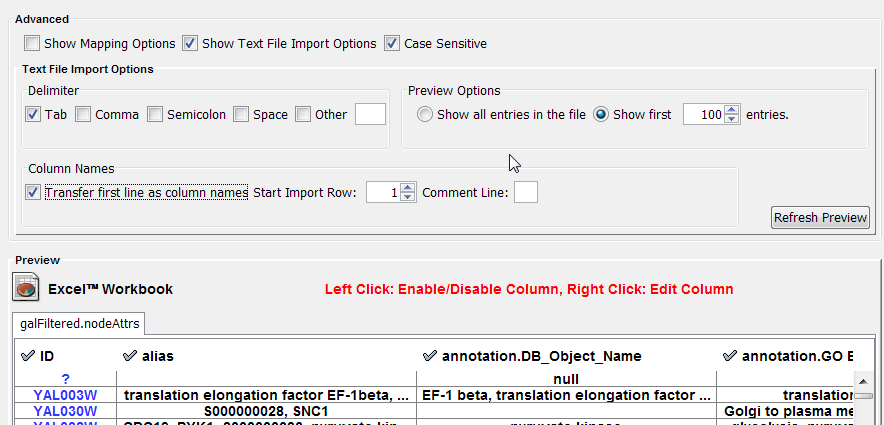
Text file import options
When the text file import options box is checked, several additional parameters can be selected to tune the way the data file will be imported. The first line can be used to specify column header names. Another data delimiter (default is tab) can be chosen, and the comment prefix (signifying lines to be ignored) can be defined.
Import from Public Databases
You can import various kinds of ID sets from the BioMart (http://www.biomart.org/index.html) public database. The BioMart web service client is integrated into Cytoscape.
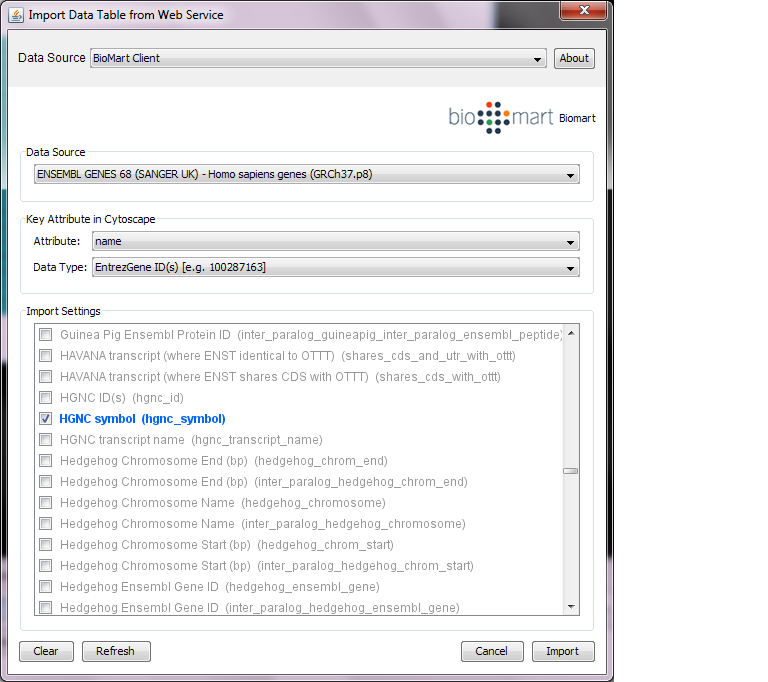
Select: File → Import → Table → Public Databases...
Select a data source. For ID mapping, select one of the Ensemble Genes data set. You need to choose the correct species for your network.
Select Attribute. If you want to import new ID sets matching current node IDs, select shared name.
Select Data Type. This should be the type of ID set selected in Attribute list. For example, if you select shared name for Attribute and your network uses Entrez Gene ID for its node ID, you need to select EntrezGene ID(s) for Data Type.
Select new ID sets from the list. Because there is a size limitation on the results returned by the!BioMart server, you can select only 3-5 attributes for each import.
Press Import.
Data in complete networks
When importing a network from a table, columns other than the node identifier can be imported as data also. For more detail on these options, please see the "Import Free-Format Table Files" section of the user manual in the Creating Networks chapter.
Table Panel
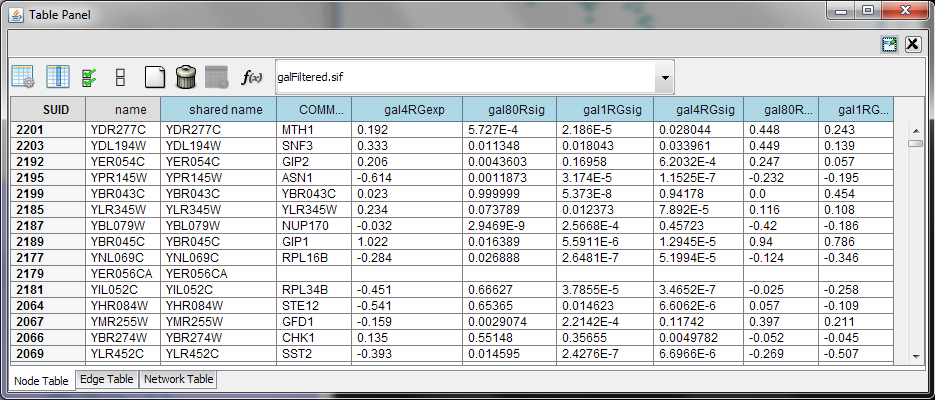
When Cytoscape is started, the Table Panel appears in the bottom CytoPanel. This panel can be hidden and restored using the View → Show/Hide Table Panel menu option. Like other CytoPanels, the panel can be undocked by pressing the little icon in the panel’s top right corner.
To swap between displaying node, edge, and network tables use the tabs on the bottom of the panel labeled "Node Table", "Edge Table", and "Network Table".
In Cytoscape 3.0 there are two display modes for the table: show selected nodes/edges only and show all rows. This configuration can be set using button (the left most) in the figure. The Table Panel displays data belonging to the currently selected network.
Using the three buttons (left 2nd to 4th) in the figure, it is possible to make some or all columns visible and hide others or all of them. Also, a new column can be created by pressing button the (left 5th) or mutable columns can be deleted by button (left 6th). Button f(x) is for writing equations to manipulate the data which is further explained in the section attribute functions and equations.
Most data values can be edited by double-clicking on their table cell (only the SUID cannot be edited). Newline characters can be inserted into String attributes either by pressing Enter or by typing "\n". Once finished editing, click outside of the editing cell in the Table Panel or press Shift-Enter to save your edits. Pressing Esc while editing will undo any changes.
Table rows in the browser can be sorted by a specific column by clicking on a column heading. A new column can be created using the Create New Column button (left 5th), and must be one of four types – integer, string, real number (floating point), or boolean. Attributes can be deleted using the Delete Attributes button (left 6th, trash can icon). NOTE: Deleting attributes removes them from Cytoscape, not just the attribute browser! To remove attributes from the browser without deleting them, simply unselect the attribute using the Select Column button (left 3rd). The right-click menu on the Table Panel has several functions, such as exporting attribute information to spreadsheet applications. For example, use the right-click menu to select the data and copy for use in a spreadsheet application.
Another type of data that can be associated with network elements are ontologies. An ontology consists of an organized set of controlled vocabulary terms that annotate the objects. Most ontologies in science are organized in a hierarchical way. In biology for example, using the Gene Ontology, the Saccharomyces Cerevisiae Cytoscape can use this ontology DAG (Directed Acyclic Graph) to annotate objects in networks. The Ontology Server is a Cytoscape feature which allows you to load, navigate, and associate ontology terms to nodes and edges in a network. Cytoscape has an GUI for loading ontology and associating it with the network elements, enabling you to load both local and remote files.
The standard file formats used in Cytoscape Ontology Server are OBO and Gene Association. The GO website details these file formats: Ontologies and Definitions: http://www.geneontology.org/GO.downloads.shtml#ont Current Associtations: http://www.geneontology.org/GO.current.annotations.shtml
An OBO file is the ontology DAG itself. This file defines the relationships between ontology terms. Cytoscape can load all ontology files written in OBO format. The full listing of ontology files are available from the Open Biomedical Ontologies (OBO) website: OBO Ontology Browser: http://obo.sourceforge.net/browse.html Sample OBO File - gene_ontology.obo: http://www.geneontology.org/ontology/gene_ontology_edit.obo
Cytoscape provides a list of ontologies available in OBO format. If an Internet connection is available, Cytoscape will import ontology and associatation files directly from the remote source. The table below lists the included ontologies. Ontology Name Description Gene Ontology Full This data source contains a full-size GO DAG, which contains all GO terms. This OBO file is written in version 1.2 format. Generic GO slim A subset of general GO Terms, including higer-level terms only. Yeast GO slim A subset of GO Terms for annotating Yeast data sets maintained by SGD. Molecule role (INOH Protein name/family name ontology) A structured controlled vocabulary of concrete and abstract (generic) protein names. This ontology is a INOH pathway annotation ontology, one of a set of ontologies intended to be used in pathway data annotation to ease data integration. This ontology is used to annotate protein names, protein family names, and generic/concrete protein names in the INOH pathway data. INOH is part of the BioPAX working group. Event (INOH pathway ontology) A structured controlled vocabulary of pathway-centric biological processes. This ontology is a INOH pathway annotation ontology, one of a set of ontologies intended to be used in pathway data annotation to ease data integration. This ontology is used to annotate biological processes, pathways, and sub-pathways in the INOH pathway data. INOH is part of the BioPAX working group. Protein-protein interaction A structured controlled vocabulary for the annotation of experiments concerned with protein-protein interactions. Pathway Ontology The Pathway Ontology is a controlled vocabulary for pathways that provides standard terms for the annotation of gene products. PATO PATO is an ontology of phenotypic qualities, intended for use in a number of applications, primarily phenotype annotation. For more information, please visit the PATO wiki (http://www.bioontology.org/wiki/index.php/PATO:Main_Page). Mouse pathology The Mouse Pathology Ontology (MPATH) is an ontology for mutant mouse pathology. This is Version 1. Human disease This ontology is a comprehensive hierarchical controlled vocabulary for human disease representation. For more information, please visit the Disease Ontology website (http://diseaseontology.sourceforge.net/). Although Cytoscape can import all kinds of ontologies in OBO format, association files are associated with specific ontologies. Therefore, you need to provide the correct ontology-specific association file to annotate nodes/edges/networks in Cytoscape. For example, while you can annotate human network data using the GO Full ontology with human Gene Association files, you cannot use a combination of the human Disease Ontology file and human Gene Association files, because the Gene Association file is only compatible with GO.
Relationships between ontology terms are usually represented as Directed Acyclic Graphs (DAGs). This is a special case of a network (or graph), where nodes are ontology terms and edges are relationships between terms. Ontology data is stored in the same data structure as normal networks. This enables users and App writers to visualize, browse and manipulate ontology DAGs just like other networks. The following is an example of visualization of an ontology DAG (Generic GO Slim): Every ontology term and relationship can have attributes. In the OBO files, ontology terms have optional fields such as definition, synonyms, comments, or cross-references. These fields will be imported as node attributes. To browse those attributes, please use the attribute browser (see the example below):
The Gene Association (GA) file provides annotation only for the Gene Ontology. It is a species-specific association file for GO terms. Sample Gene Association File (gene_association.sgd - association file for yeast): If you have a network file and an association file, they should have a common key to map attributes onto network data. If those two do not have a common key, you need to do an extra step to add a shared key by mapping the current key to a common key as described above (Node Name Mapping).
For convenience, Cytoscape has a list of URLs for commonly used ontology data and a complete set of Gene Association files. To import Gene Ontology and Gene Association files for the loaded networks, please follow these steps: Important: All data sources in the preset list are remote URLs, meaning a network connection is required!
Select File → Import → Ontology and Annotation... to open the "Import Ontology and Annotation" window. From the Annotation dropdown list, select a gene association file for your network. For example, if you want to annotate the yeast network, select "Gene Association file for Saccharomyces cerevisiae".
Ontologies
GO 8150 biological_process
GO 7582 physiological processes
GO 8152 metabolism
GO 44238 primary metabolism
GO 19538 protein metabolism
GO 6412 protein biosynthesisGraphical View of GO Term 6412: protein biosynthesis 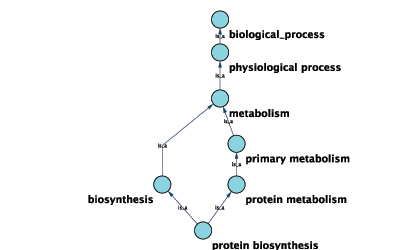
Ontology and Association File Format
OBO File
format-version: 1.2
date: 27:11:2006 17:12
saved-by: midori
auto-generated-by: OBO-Edit 1.002
subsetdef: goslim_generic "Generic GO slim"
subsetdef: goslim_goa "GOA and proteome slim"
subsetdef: goslim_plant "Plant GO slim"
subsetdef: goslim_yeast "Yeast GO slim"
subsetdef: gosubset_prok "Prokaryotic GO subset"
default-namespace: gene_ontology
remark: cvs version: $Revision: 5.49 $
[Term]
id: GO:0000001
name: mitochondrion inheritance
namespace: biological_process
def: "The distribution of mitochondria, including the mitochondrial genome, into daughter cells after mitosis or meiosis, mediated by interactions between mitochondria and the cytoskeleton." [GOC:mcc, PMID:10873824, PMID:11389764]
synonym: "mitochondrial inheritance" EXACT []
is_a: GO:0048308 ! organelle inheritance
is_a: GO:0048311 ! mitochondrion distribution
[Term]
id: GO:0000002
name: mitochondrial genome maintenance
namespace: biological_process
def: "The maintenance of the structure and integrity of the mitochondrial genome." [GOC:ai]
is_a: GO:0007005 ! mitochondrion organization and biogenesis
Default List of Ontologies
Visualize and Browse Ontology DAG (for Advanced Users)
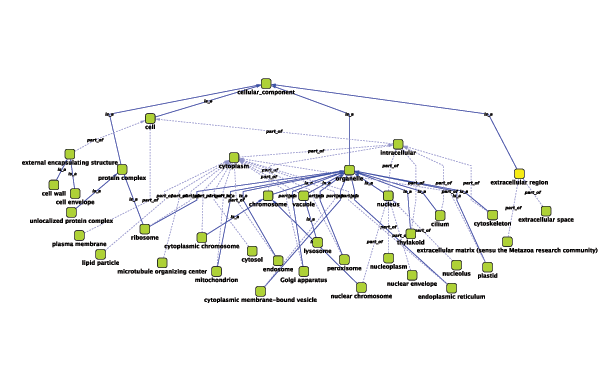
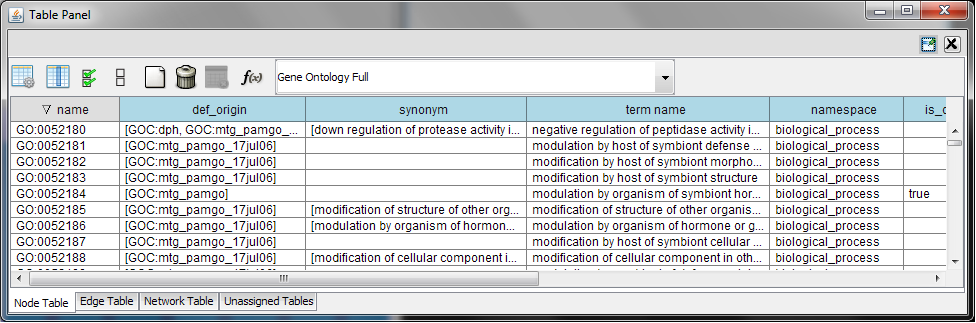
Gene Association File
SGD S000003916 AAD10 GO:0006081 SGD_REF:S000042151|PMID:10572264 ISS P aryl-alcohol dehydrogenase (putative) YJR155W gene taxon:4932 20020902 SGD
SGD S000005275 AAD14 GO:0008372 SGD_REF:S000069584 ND C aryl-alcohol dehydrogenase (putative) YNL331C gene taxon:4932 20010119 SGD
Import Gene Ontology and Gene Association Files
Step 1. Select an Annotation File
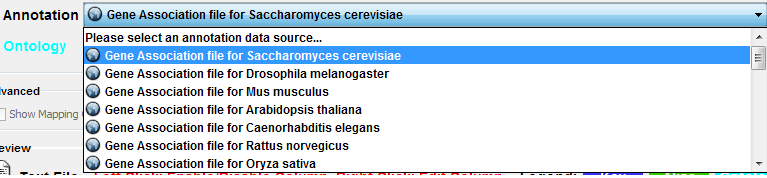
Step 2. Select an Ontology File
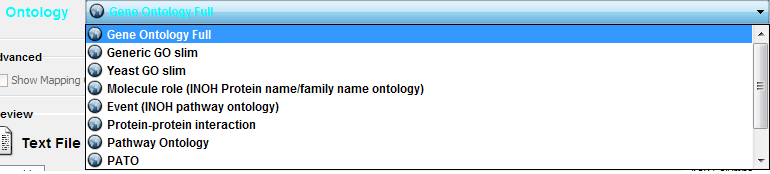
Step 3. Import the files
Navigation and Layout
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
Basic Network Navigation
Cytoscape uses a Zoomable User Interface for navigating and viewing networks. ZUIs use two mechanisms for navigation: zooming and panning. Zooming increases or decreases the magnification of a view based on how much or how little a user wants to see. Panning allows users to move the focus of a screen to different parts of a view.
Zoom
Cytoscape provides two mechanisms for zooming: toolbar buttons and the scroll wheel. Use the zooming buttons located on the toolbar to zoom in and out of the interaction network shown in the current network display. Zoom icons are detailed below:

From Left to Right:
- Zoom In
- Zoom Out
- Zoom Out to Display all of Current Network
- Zoom Selected Region
Using the scroll wheel, you can zoom in by scrolling up and zoom out by scrolling downwards. These directions are reversed on Macs with natural scrolling enabled (the default for Mac OS X Lion and newer versions).
Pan
There are several ways to pan the network:
- Middle Click and Drag - You can pan the network image by holding down the middle mouse button and moving the mouse.
- Command Key + Drag (Mac only) - If you use Mac without middle button, you can pan the view by holding down Command key and then drag.
Dragging Box on Network Overview - You can also pan the image by holding down the left mouse button over the blue box in the overview panel in the lower part of the Network tab in the Control Panel.
Other Mouse Behaviors
Select
Click the left mouse button on a node or edge to select that object. You can hold down the Shift key (or Ctrl key on PCs) to select more than one node/edge or you can hold down the left mouse button and drag the mouse to select groups of nodes/edges.
Context
Click the right mouse button (or Ctrl+left mouse button on Macs) on a node/edge to launch a context-sensitive menu with additional information about the node/edge.
Node Context Menu
This menu can change based on the current context. For nodes, it typically shows:
- Add
- Edit
- Select
- Group
- Nested Networks
- Apps
- External Links
- Preferences
Edges usually have the following menu:
- Edit
- Select
- Apps
- External Links
- Preferences
Apps can contribute their own items into node and edge context menus. These additions usually appear in the Apps section of the context menu.
Nested Network Node Context Menu
Add Nested Network: Lets the user select any network in Cytoscape as the current node's nested network. If the current node already has a nested network it will be replaced.
Remove Nested Network: Removes the currently associated nested network from a node. The associated network is not deleted. Only the association between the node and the network is removed.
Go to Nested Network: The current node's nested network will be the current network view and have the focus. Should a network view for the nested network not exist, it will be created.
More information about nested networks can be found in the Nested Networks section.
Automatic Layout Algorithms
The Layout menu has an array of features for organizing the network visually according to one of several algorithms, aligning and rotating groups of nodes, and adjusting the size of the network. Cytoscape layouts have three different sources, which are reflected in the Layout menu.
With the exception of the yFiles layouts (explained below), Cytoscape Layouts have the option to operate on only the selected nodes, and all provide a Settings... panel to change the parameters of the algorithm. Most of the Cytoscape layouts also partition the graph before performing the layout. In addition, many of these layouts include the option to take either node or edge columns into account. A few of the layout algorithms are:
Grid Layout
The grid layout is a simple layout the arranges all of the nodes in a square grid. This is the default layout and is always available as part of the Cytoscape core. It is available by selecting Layout → Grid Layout. A sample screen shot is shown above.
Edge-weighted Spring-Embedded Layout
The spring-embedded layout is based on a “force-directed” paradigm as implemented by Kamada and Kawai (1988). Network nodes are treated like physical objects that repel each other, such as electrons. The connections between nodes are treated like metal springs attached to the pair of nodes. These springs repel or attract their end points according to a force function. The layout algorithm sets the positions of the nodes in a way that minimizes the sum of forces in the network. This algorithm can be applied to the entire network or a portion of it by selecting the appropriate options from Layout → Edge-weighted Spring Embedded.
Attribute Circle Layout
The Attribute Circle layout is a quick, useful layout, particularly for small networks, that will locate all of the nodes in the network around a circle. The node order is determined by a user-selected node column. The result is that all nodes with the same value for that column are located together around the circle. Using Layout → Attribute Circle Layout → column to put all nodes around a circle using column to position them. The sample screen shot above shows the a subset of the galFiltered network organized by node degree.
Group Attributes Layout
The Group Attributes layout is similar to the Attribute Circle layout described above except that instead of a single circle with all of the nodes, each set of nodes that share the same value for the column are laid out in a separate circle. The same network shown above (network generated by PSICQUIC Client) is shown above, using Layout → Group Attributes Layout → taxonomy.
Prefuse Force Directed Layout
The force-directed layout is a layout based on the "force-directed" paradigm. This layout is based on the algorithm implemented as part of the prefuse toolkit provided by Jeff Heer. The algorithm is very fast and with the right parameters can provide a very visually pleasing layout. The Force Directed Layout will also accept a numeric edge column to use as a weight for the length of the spring, although this will often require more use of the Settings... dialog to achieve the best layout. This algorithm is available by selecting Layout → Prefuse Force-Directed Layout → (unweighted) or the edge column you want to use as a weight. A sample screen shot showing a portion of the galFiltered network provided in sample data is provided above.
yFiles Layouts
yFiles layouts are a set of commercial layout algorithms which are provided courtesy of yWorks. Due to license restrictions, the detailed parameters for these layouts are not available (there are no yFiles entries in the Layout → Settings...). The main layout algorithms provided by yFiles are:
yFiles Organic Layout
The organic layout algorithm is a kind of spring-embedded algorithm that combines elements of the other algorithms to show the clustered structure of a graph. This algorithm is available by selecting Layout → yFiles Layouts → Organic.
yFiles Circular Layout
This algorithm produces layouts that emphasize group and tree structures within a network. It partitions the network by analyzing its connectivity structure, and arranges the partitions as separate circles. The circles themselves are arranged in a radial tree layout fashion. This algorithm is available by selecting Layout → yFiles Layouts → Circular.
yFiles Hierarchical Layout
The hierarchical layout algorithm is good for representing main direction or "flow" within a network. Nodes are placed in hierarchically arranged layers and the ordering of the nodes within each layer is chosen in such a way that minimizes the number of edge crossings. This algorithm is available by selecting Layout → yFiles Layouts → Hierarchical.
Layout Parameters
Many layouts have adjustable parameters that are exposed through the Layouts → Settings... menu option. The Layout Settings dialog, which allows you to choose which layout algorithm settings to adjust, is shown below. The settings presented vary by algorithm and only those algorithms that allow access to their parameters will appear in the drop-down menu at the top of the dialog. Once you've modified a parameter, clicking the Execute Layout button will apply the layout.
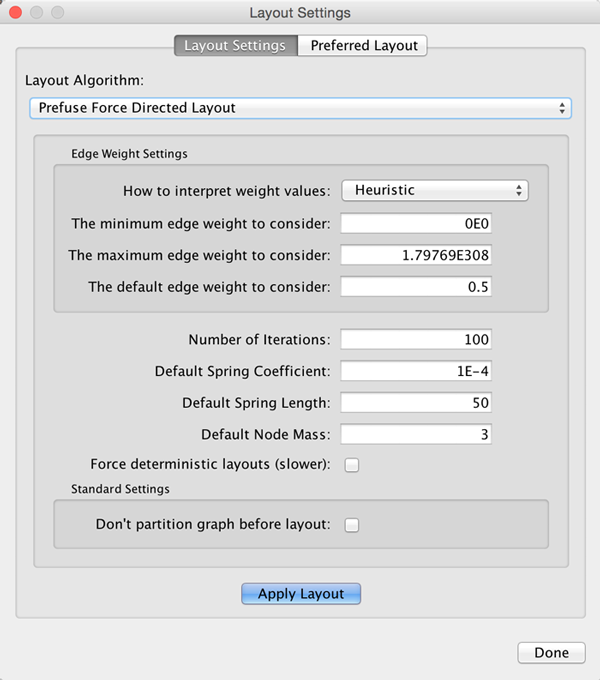
Edge Bend and Automatic Edge Bundling
From Cytoscape 3.0, Edge Bend is a regular edge property and can be used as a part of a Style. Just like any other edge property, you can select a Default Value, a Mapping and use Bypass for select nodes. In the Styles tab, select the Bend property from the Properties drop-down and click on either the Default Value, Mapping or Bypass cell to bring up the Edge Bend Editor. In the editor, you can add as many handles as you want to the edge using Alt-Click on Windows, Option-Click on Mac, or Ctrl-Alt-Click on Linux.
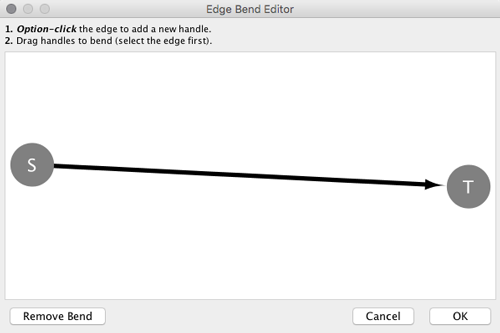
To clear all edge bends, select Layout → Clear All Edge Bends.
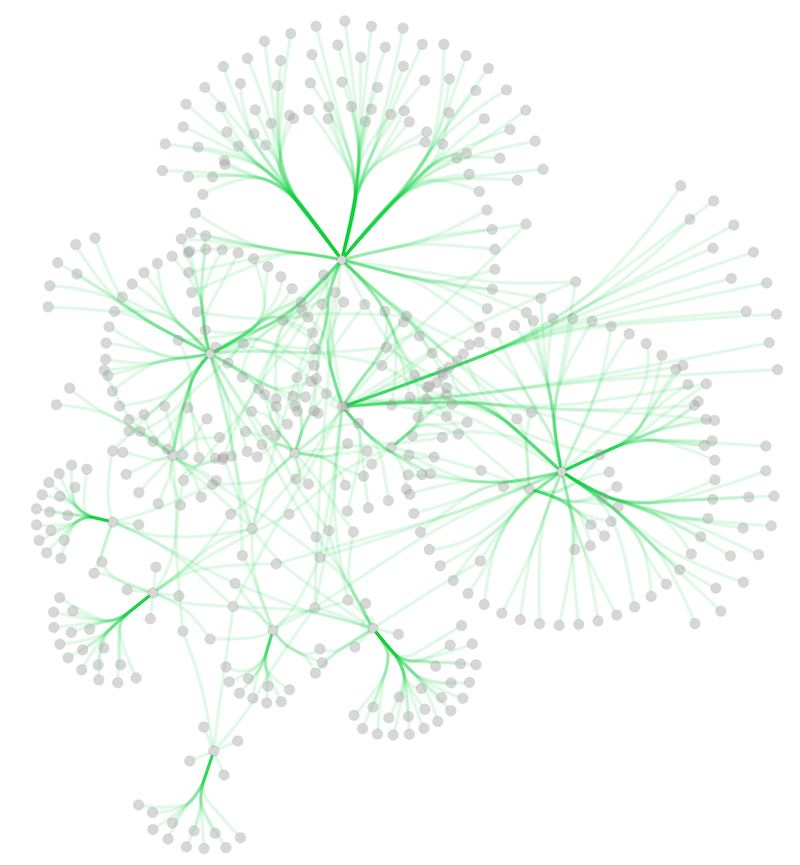
In addition to adding handles manually, you can use the Bundle Edges function to bundle all or selected edges automatically.
Select Layout → Bundle Edges → All Nodes and Edges.
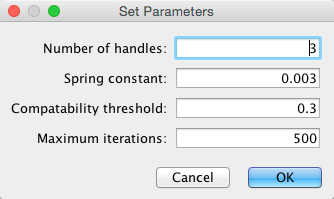
- Set parameters.
Details of the algorithm is described in this paper.
- Press OK to run. Edge bundling may take a long time if the number of edges is large.
If it takes too long, try decreasing Maximum Iterations.
For large, dense networks, try setting Maximum iterations in the range of 500 - 1000.
Note: The handle locations will be optimized for current location of nodes. If you move node positions, you need to run the function again to get proper result.
Manual Layout
The simplest method to manually organize a network is to click on a node and drag it. If you select multiple nodes, all of the selected nodes will be moved together.
Rotate
Selecting the Layout → Rotate option will show the Rotate window in the Tool Panel. This function will either rotate the entire network or a selected portion of the network. The image below shows a network with selected nodes rotated.
Before
After
Scale
Selecting the Layout → Scale option will open the Scale window in the Tool Panel. This function will scale the position of the entire network or of the selected portion of the network. Note that only the position of the nodes will be scaled, not the node sizes. Node size can be adjusted using Styles. The image below shows selected nodes scaled.
Before
After
Align, Distribute and Stack
Selecting the Layout → Align/Distribute option will open the Align and Distribute window in the Tool Panel. Align provides different options for either vertically or horizontally aligning selected nodes against a line. The differences are in what part of the node gets aligned, e.g. the center of the node, the top of the node, the left side of the node. Distribute evenly distributes selected nodes between the two most distant nodes along either the vertical or horizontal axis. The differences are again a function what part of the node is used as a reference point for the distribution. Stack vertically or horizontally stacks selected nodes with the full complement of alignment options. The table below provides a description of what each button does.
Button |
Before |
After |
Description of Align Options |
|
|
|
Vertical Align Top - The tops of the selected nodes are aligned with the top-most node. |
|
|
|
Vertical Align Center - The centers of the selected nodes are aligned along a line defined by the midpoint between the top and bottom-most nodes. |
|
|
|
Vertical Align Bottom - The bottoms of the selected nodes are aligned with the bottom-most node. |
|
|
|
Horizontal Align Left - The left hand sides of the selected nodes are aligned with the left-most node. |
|
|
|
Horizontal Align Center - The centers of the selected nodes are aligned along a line defined by the midpoint between the left and right-most nodes. |
|
|
|
Horizontal Align Right - The right hand sides of the selected nodes are aligned with the right-most node. |
Button |
Before |
After |
Description of Distribute Options |
|
|
|
Vertical Distribute Top - The tops of the selected nodes are distributed evenly between the top-most and bottom-most nodes, which should stay stationary. |
|
|
|
Vertical Distribute Center - The centers of the selected nodes are distributed evenly between the top-most and bottom-most nodes, which should stay stationary. |
|
|
|
Vertical Distribute Bottom - The bottoms of the selected nodes are distributed evenly between the top-most and bottom-most nodes, which should stay stationary. |
|
|
|
Horizontal Distribute Left - The left hand sides of the selected nodes are distributed evenly between the left-most and right-most nodes, which should stay stationary. |
|
|
|
Horizontal Distribute Center - The centers of the selected nodes are distributed evenly between the left-most and right-most nodes, which should stay stationary. |
|
|
|
Horizontal Distribute Right - The right hand sides of the selected nodes are distributed evenly between the left-most and right-most nodes, which should stay stationary. |
Button |
Before |
After |
Description of Stack Options |
|
|
|
Vertical Stack Left - Vertically stacked below top-most node with the left-hand sides of the selected nodes aligned. |
|
|
|
Vertical Stack Center - Vertically stacked below top-most node with the centers of selected nodes aligned. |
|
|
|
Vertical Stack Right - Vertically stacked below top-most node with the right-hand sides of the selected nodes aligned. |
|
|
|
Horizontal Stack Top - Horizontally stacked to the right of the left-most node with the tops of the selected nodes aligned. |
|
|
|
Horizontal Stack Center - Horizontally stacked to the right of the left-most node with the centers of selected nodes aligned. |
|
|
|
Horizontal Stack Bottom - Horizontal Stack Center - Horizontally stacked to the right of the left-most node with the bottoms of the selected nodes aligned. |
Node Movement and Placement
In addition to the ability to click on a node and drag it to a new position, Cytoscape now has the ability to move nodes using the arrow keys on the keyboard. By selecting one or more nodes using the mouse and clicking one of the arrow keys (←, →, ↑, ↓) the selected nodes will move one pixel in the chosen direction. If an arrow key is pressed while holding the Shift key down, the selected nodes will 15 pixels in the chosen direction.
Finding and Filtering Nodes and Edges
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
Search Bar
You can search for nodes and edges by column value directly through Cytoscape's tool bar. For example, to select nodes or edges with a column value that starts with "STE", type ste* in the search bar. The search is case-insensitive. The * is a wildcard character that matches zero or more characters, while "?" matches exactly one character. So ste? would match "STE2" but would not match "STE12". Searching for ste* would match both.
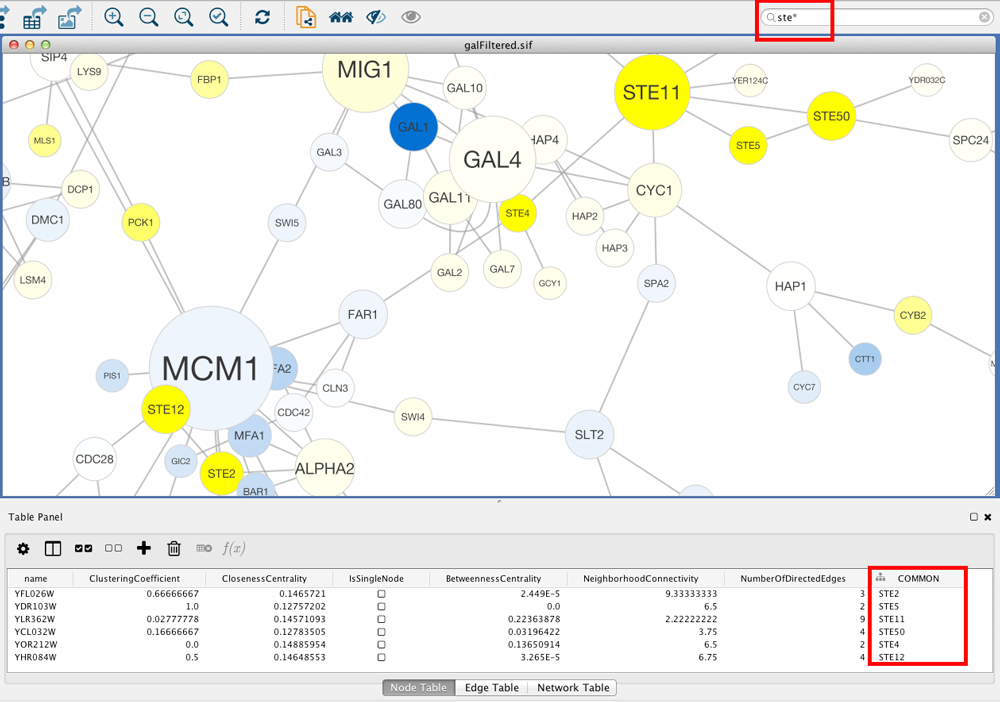
To search a specific column, you can prefix your search term with the column name followed by a :. For example, to select nodes and edges that have a "COMMON" column value that starts with "STE", use common:ste*. If you don't specify a particular column, all columns will be searched.
Columns with names that contain spaces, quotes, or characters other than letters and numbers currently do not work when searching a specific column. This will be fixed in a future release.
To search for column values that contain special characters you need to escape those characters using a "\". For example, to search for "GO:1232", use the query GO\:1232. The complete list of special characters is:
+ - & | ! ( ) { } [ ] ^ " ~ * ? : \Note: Escaping characters only works when searching all columns. It currently does not work for column-specific searching. This will be fixed in a future release.
Filters
Cytoscape 3 provides a new user interface for filtering nodes and edges. These tools can be found in the Select panel:
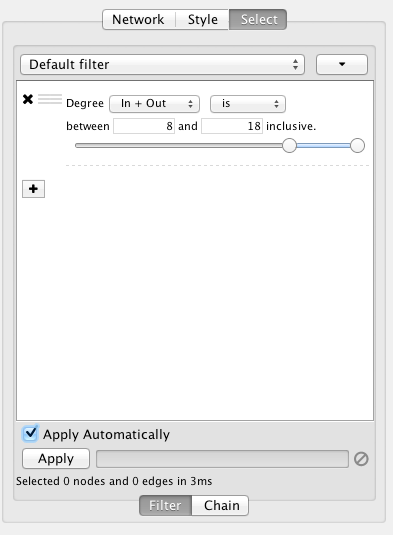
There are two types of filters. On the Filter tab are narrowing filters, which can be combined into a tree. On the Chain tab are chainable filters, which can be combined in a linear chain.
Narrowing Filters
Narrowing filters are applied to the entire network, and are used to select a subset of nodes or edges in a network based on user-specified constraints. For example, you can find edges with a weight between 0 and 5.5, or nodes with degree less than 3. A filter can contain an arbitrary number of sub-filters.
To add a filter click on the "+" button. To delete a filter (and all its sub-filters) click the "x" button. To move a filter grab the handle  with the mouse and drag and drop the filter on its intended destination. Dropping a filter on top of another filter will group the filters into a composite filter.
with the mouse and drag and drop the filter on its intended destination. Dropping a filter on top of another filter will group the filters into a composite filter.
Interactive Filter Application Mode
Due to the nature of narrowing filters, Cytoscape can apply them to a network efficiently and interactively. Some filters even provide slider controls to quickly explore different thresholds. This is the default behavior on smaller networks. For larger networks, Cytoscape automatically disables this interactivity. You can override this by manually checking the Apply Automatically box above the Apply button:
Cytoscape comes packaged with the following narrowing filters:
Column Filter
This filter will match nodes or edges that have particular column values. For numeric columns sliders are provided to set minimum and maximum values, or the values may be entered manually.
From string columns, a variety of matching options are provided:
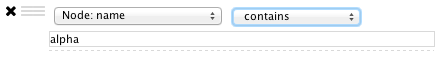
For example, column values can be checked to see if they contain or match exactly the text entered in the text box. More complex matching criteria can be specified by using a Java-style regular expression.
By default string matching is case insensitive. Case sensitive matching requires the use of a regular expression that starts with "(?-i)". For example to match the text "ABC" in a case sensitive way use the following regular expression: "(?-i)ABC".
Cytoscape uses Java regular expression syntax.
Degree Filter
The degree filter matches nodes with a degree that falls within the given minimum and maximum values, inclusive. You can choose whether the filter operates on the in-degree, out-degree or overall (in + out) degree.
Topology Filter
The topology filter matches nodes having a certain number of neighbors which are within a fixed distance away, and which match a sub-filter. The thresholds for the neighborhood size and distance can be set independently, and the sub-filter is applied to each such neighbor node.
The topology filter will successfully match a node if the sub-filter matches against the required number of neighbor nodes.
Grouping and Organizing Filters
By default, nodes and edges need to satisfy the constraints of all your filters. You can change this so that instead, only the constraints of at least one filter needs to be met in order to match a node or edge. This behavior is controlled by the Match all/any drop-down box. This appears once your filter has more than one sub-filter. For example, suppose you wanted to match nodes with column COMMON containing ste or cdc, but you only want nodes with degree 5 or more, you'd first construct a filter that looks like this:
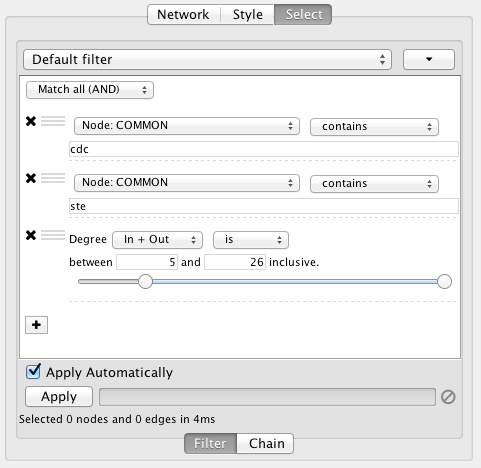
This filter will match nodes where COMMON contains ste and cdc. To change this to a logical or operation, drag either of the column filters by its handle  onto the other column filter to create a new group. Now change the group's matching behavior to Match any:
onto the other column filter to create a new group. Now change the group's matching behavior to Match any:
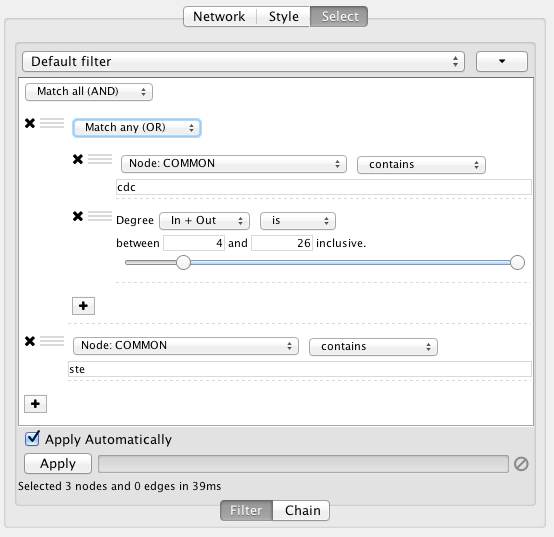
You can also reorder filters by dropping them in-between existing filters.
Chainable Filters
Chainable filters are combined in an ordered list. The nodes and edges in the output of a filter become the input of the next filter in the chain. The first filter in the chain gets its input from the current selection or from a filter on the Filter tab. The output of the last filter becomes the new selection.
You can specify the input to the first filter in the chain by selecting a Start with, where Current selection refers to the nodes and edges currently selected. You can also choose a narrowing filter, which produces a different set of selected nodes and edges.
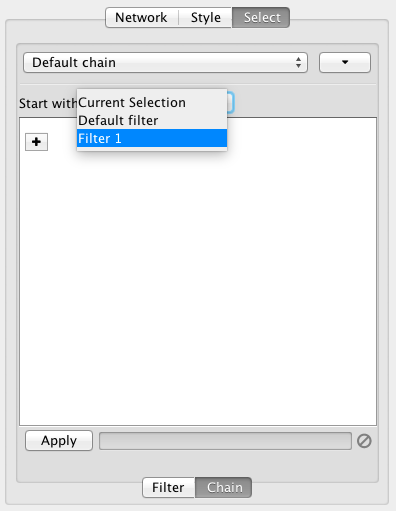
Chainable filters can be reordered by dragging one by the handle and dropping it between existing filters.
Cytoscape currently bundles the following chainable filters:
Edge Interaction Transformer
This transformer will go through all the input edges and selectively add their source nodes, target nodes, or both, to the output. This is useful for adding nodes that are connected to edges that match a particular filter.
Output options:
- Add (default): Automatically includes all input nodes and edges in the output, and adds source or target nodes from input edges to the output.
- Replace with: Does not automatically include input nodes and edges in the output. Only outputs nodes that match the filter.
A sub-filter may be added as well. When a sub-filter is present the source/target nodes must match the filter to be included in the output.
Node Adjacency Transformer
This transformer is used to add nodes and edges that are adjacent to the input nodes. A sub-filter may be specified as well.
Note that pressing the Apply button repeatedly may cause the selection to continuously expand. This allows adjacent nodes that are at greater distances to be added.
Output options:
- Add (default): Automatically includes all input nodes and edges in the output, and adds selected adjacent nodes and edges.
- Replace with: Only outputs the adjacent nodes/edges.
Select options:
- Adjacent nodes: Output nodes that are adjacent to the input nodes.
- Adjacent edges: Output edges that are adjacent (incident) to the input nodes.
- Adjacent nodes and edges (default): Output both nodes and edges that are adjacent to the input nodes.
Edge direction options. (Hidden by default, click the small arrow icon to reveal.):
- Incoming: Only include adjacent nodes/edges when the adjacent edge is incoming.
- Outgoing: Only include adjacent nodes/edges when the adjacent edge is outgoing.
- Incoming and Outgoing (default): Ignore the directionality of adjacent edges.
Sub-filter options. (Available when a sub-filter has been added.):
- Adjacent nodes (default): The sub-filter is only applied to adjacent nodes. (Edges to the adjacent nodes are still included in the output.)
- Adjacent edges: The sub-filter is only applied to adjacent edges. (Nodes connected to the adjacent edges are still included in the output.)
- Adjacent nodes and edges: Both the adjacent edge and its connected node must match the filter. Note that for a filter to match an edge and a node at the same time it should be a compound filter that is set to "Match any (OR)".
Working with Narrowing and Chainable Filters
The name of active filter appears in the drop-down box at the top of Select panel. Beside this is the options button which will allow you to rename, remove or export the active filter. It also lets you create a new filter, or import filters.
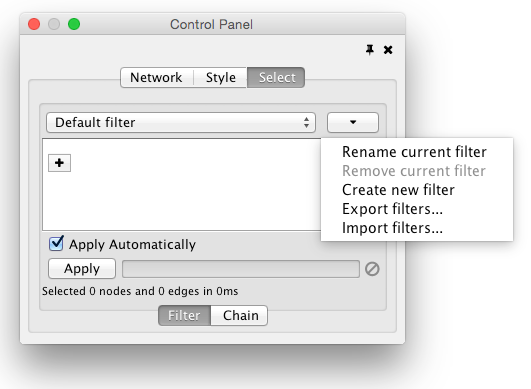
At the bottom of the Select panel, there is an Apply button that will re-apply the active filter. On the opposite side of the progress bar is the cancel button, which will let you interrupt a long-running filter.
The Select Menu
The Select → Nodes and Select → Edges menus provide several mechanisms for selecting nodes and edges. Most options are fairly straightforward; however, some need extra explanation.
Select → Nodes → From ID List File... selects nodes based on node identifiers found in a specified file. The file format is simply one node id per line:
Node1 Node2 Node3 ...
Editing Networks
Using Cytoscape you can edit networks, add nodes and edges. In Cytoscape 3, editing can be done using right click menus on the network view panel. To start editing a new network, create a new network by going to File → New → Network → Empty Network.
Adding Node
To add a new node, right click on an empty space of the network view panel. Select Add → Node item from the pop-up menu. Image below shows the right click menu for creating a new node.
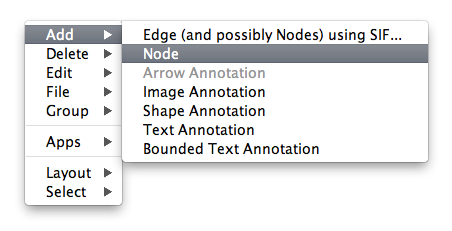
Adding Edge
For adding an edge to connect to nodes, right-click on the source node. Select Edit → Add Edge from the pop-up menu. Next, move pointer and click on the target node. Images below show the two steps for drawing an edge between two nodes. You can abort the drawing of the edge by pressing Esc key.
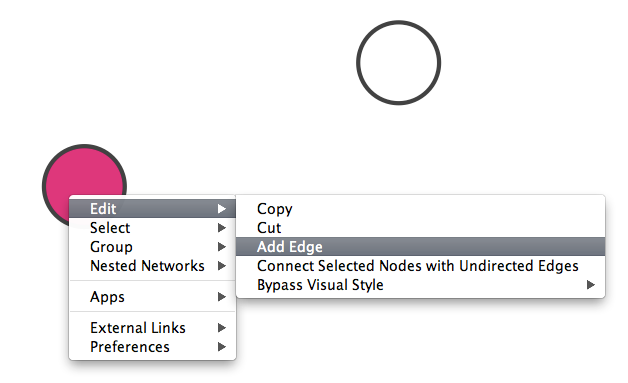
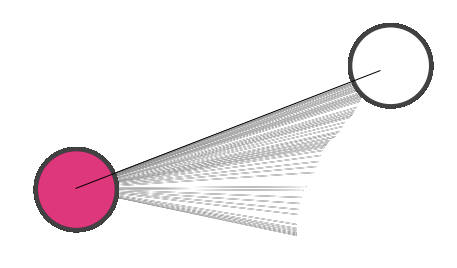
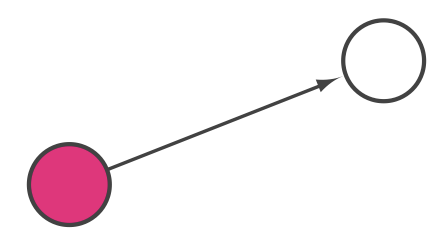
You can delete nodes and edges by selecting a number of nodes and edges, then selecting Edit → Delete Selected Nodes and Edges. You can recover any nodes and edges deleted from a network by going to Edit → Undo.
Nested Networks
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
Creating Nested Networks
There are currently two ways in which a user can create nested networks.
By importing a Nested Network Format (NNF) file. (See: NNF Network Format).
By manually constructing networks and assigning nested networks to individual nodes through the right-click node context menu. (See Nested Network Node Context Menu).
Visualization of Nested Networks
Nodes containing nested networks that are zoomed in sufficiently display an image for the nested network. If no current network view exists for the nested network the image will be a default icon, otherwise it will be a low-resolution rendering of the nested network's current layout.
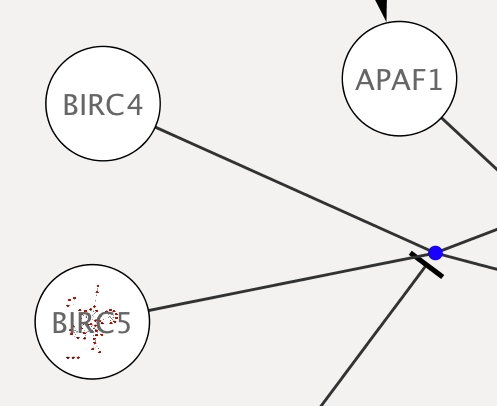
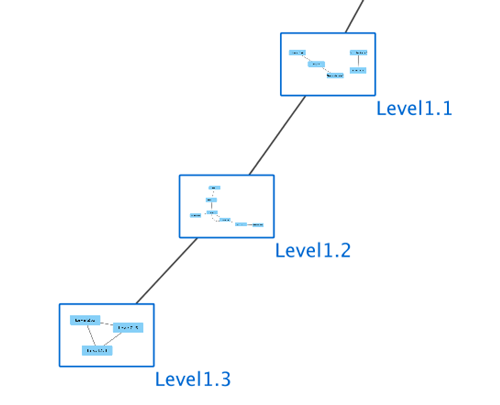
Programmatically Manipulating Nested Networks
The giny.model.node interface defines two methods:
public void setNestedNetwork(final GraphPerspective graphPerspective); This will assign a "network" (GraphPerspective) to a Node or replace the assigned network at a node if there was a prior existing one. In order to remove a nested network, please pass null to this method.
public GraphPerspective getNestedNetwork(); If a Node has an associated nested network, it will be returned by this method. If no associated nested network exists, null will be returned instead.
Rendering Engine
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
What is Level of Detail (LOD)?
Cytoscape 3.0 retains the rendering engine found in version 2.8. It is to be able to display large networks (>10,000 nodes), yet retain interactive speed. To accomplish this goal, a technique involving level of detail (LOD) is being used. Based on the number of objects (nodes and edges) being rendered, an appropriate level of detail is chosen. For example, by default, node labels (if present) are only rendered when less than 200 nodes are visible because drawing text is a relatively expensive operation. This can create some unusual behavior. If the screen currently contains 198 nodes, node labels will be displayed. If you pan across the network, such that now 201 nodes are displayed, the node labels will disappear. As another example, if the sum of rendered edges and rendered nodes is greater than or equal to a default value of 4000, a very coarse level of detail is chosen, where edges are always straight lines, nodes are always rectangles, and no anti-aliasing is done. The default values used to determine these thresholds can be changed by setting properties under Edit → Preferences → Properties....
Low LOD vs High LOD
Network with Low LOD |
Network with High LOD |
|
|
With low LOD values, all nodes are displayed as square and anti-aliasing is turned off. With high LOD values, anti-aliasing is turned on and nodes are displayed as actual shape user specified in the Style.
Parameters for Controlling LOD
NOTE: The greater these thresholds become, the slower performance will become. If you work with small networks (a few hundred nodes), this shouldn't be a problem, but for large networks it will produce noticeable slowing. The various thresholds are described below.
render.coarseDetailThreshold |
If the sum of rendered nodes and rendered edges equals to or exceeds this number, a very coarse level of detail will be chosen and all other detail parameters will be ignored. If the total number of nodes and edges is below this threshold, anti-alias will be turned on; this value defaults to 4000. |
|
render.nodeBorderThreshold |
If the number of rendered nodes equals to or exceeds this number, node borders will not be rendered; this value defaults to 400. |
|
render.nodeLabelThreshold |
If the number of rendered nodes equals to or exceeds this number, node labels will not be rendered; this value defaults to 200. |
|
render.edgeArrowThreshold |
If the number of rendered edges equals to or exceeds this number, edge arrows will not be rendered; this value defaults to 600. |
|
render.edgeLabelThreshold |
If the number of rendered edges equals to or exceeds this number, edge labels will not be rendered; this value defaults to 200. |
|
When printing networks or exporting to formats such as PostScript, the highest level of detail is always chosen, regardless of what is currently being displayed on the screen.
Force to Display Detail
If you want to display every detail of the network regardless of LOD values, you can toggle to full details mode by View → Show Graphics Details (or CTR+SHIFT+F on Windows/Linux, Command+SHIFT+F for Mac). This option forces the display of all graphics details. If the network is large, this option slows down rendering speed. To hide details, select the menu item again (View → Hide Graphics Details).
Annotation
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
GO 8150 biological_process
- GO 7582 physiological processes
- GO 8152 metabolism
- GO 44238 primary metabolism
- GO 19538 protein metabolism
- GO 6412 protein biosynthesis
- GO 19538 protein metabolism
- GO 44238 primary metabolism
- GO 8152 metabolism
Graphical View of GO Term 6412: protein biosynthesis
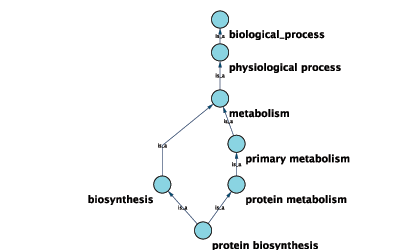
Cytoscape can use this ontology DAG (Directed Acyclic Graph) to annotate objects in networks. The Ontology Server (originally called "BioDataServer") is a Cytoscape feature which allows you to load, navigate, and assign annotation terms to nodes and edges in a network. Cytoscape 2.4 now has an enhanced GUI for loading ontology and associated annotation, enabling you to load both local and remote files.
Ontology and Annotation File Format
The standard file formats used in the Cytoscape Ontology Server are OBO and Gene Association. The GO website details these file formats:
Ontologies and Definitions: http://www.geneontology.org/GO.downloads.shtml#ont
Current Annotations: http://www.geneontology.org/GO.current.annotations.shtml
Default List of Ontologies
Cytoscape provides a list of ontologies available in OBO format. If an Internet connection is available, Cytoscape will import ontology and annotation files directly from the remote source. The table below lists the included ontologies.
Ontology Name |
Description |
Gene Ontology Full |
This data source contains a full-size GO DAG, which contains all GO terms. This OBO file is written in version 1.2 format. |
Generic GO slim |
A subset of general GO Terms, including higer-level terms only. |
Yeast GO slim |
A subset of GO Terms for annotating Yeast data sets maintained by SGD. |
Molecule role (INOH Protein name/family name ontology) |
A structured controlled vocabulary of concrete and abstract (generic) protein names. This ontology is a INOH pathway annotation ontology, one of a set of ontologies intended to be used in pathway data annotation to ease data integration. This ontology is used to annotate protein names, protein family names, and generic/concrete protein names in the INOH pathway data. INOH is part of the BioPAX working group. |
Event (INOH pathway ontology) |
A structured controlled vocabulary of pathway-centric biological processes. This ontology is a INOH pathway annotation ontology, one of a set of ontologies intended to be used in pathway data annotation to ease data integration. This ontology is used to annotate biological processes, pathways, and sub-pathways in the INOH pathway data. INOH is part of the BioPAX working group. |
Protein-protein interaction |
A structured controlled vocabulary for the annotation of experiments concerned with protein-protein interactions. |
Pathway Ontology |
The Pathway Ontology is a controlled vocabulary for pathways that provides standard terms for the annotation of gene products. |
PATO |
PATO is an ontology of phenotypic qualities, intended for use in a number of applications, primarily phenotype annotation. For more information, please visit the PATO wiki (http://www.bioontology.org/wiki/index.php/PATO:Main_Page). |
Mouse pathology |
The Mouse Pathology Ontology (MPATH) is an ontology for mutant mouse pathology. This is Version 1. |
Human disease |
This ontology is a comprehensive hierarchical controlled vocabulary for human disease representation. For more information, please visit the Disease Ontology website (http://diseaseontology.sourceforge.net/). |
Although Cytoscape can import all kinds of ontologies in OBO format, annotation files are associated with specific ontologies. Therefore, you need to provide the correct ontology-specific annotation file to annotate nodes/edges/networks in Cytoscape. For example, while you can annotate human network data using the GO Full ontology with human Gene Association files, you cannot use a combination of the human Disease Ontology file and human Gene Association files, because the Gene Association file is only compatible with GO.
Gene Association File
The Gene Association files provide annotation only for the Gene Ontology. It is a species-specific annotation file for GO terms. Gene Association files will only work with Gene Ontology annotation.
Sample Gene Association File (gene_association.sgd - annotation file for yeast):
SGD |
S000003916 |
AAD10 |
GO:0006081 |
SGD_REF:S000042151|PMID:10572264 |
ISS |
P |
aryl-alcohol dehydrogenase (putative) |
YJR155W gene |
taxon:4932 |
SGD |
S000005275 |
AAD14 |
GO:0008372 |
SGD_REF:S000069584 |
ND |
C |
aryl-alcohol dehydrogenase (putative) |
YNL331C gene |
taxon:4932 |
Import Ontology and Annotation
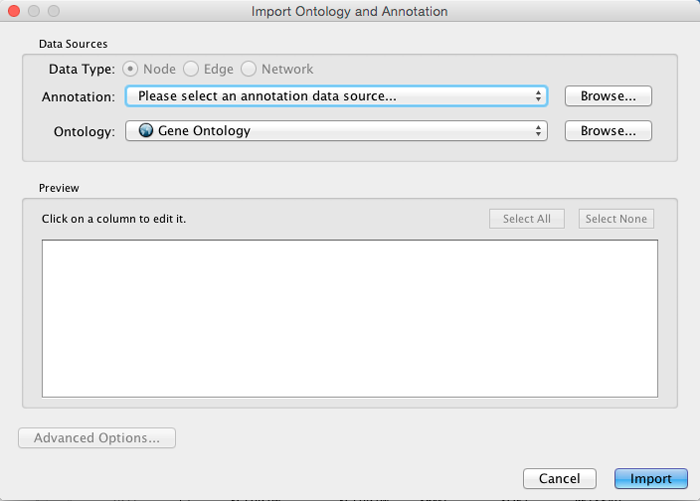
Cytoscape provides a graphical user interface to import both ontology and annotation files at the same time.
Note: All data sources in the preset list are remote URLs, meaning a network connection is required.
Select File → Import → Ontology and Annotation... to open the "Import Ontology and Annotation" interface. From the Annotation drop-down list, select a gene association file for your network. For example, if you want to annotate the yeast network, select "Gene Association file for Saccharomyces cerevisiae".
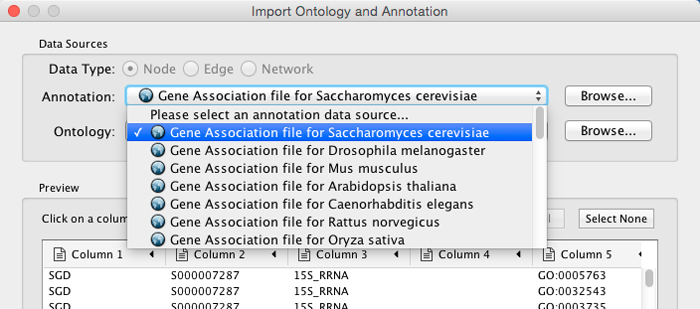
* Select an Ontology data (OBO file) from the Ontology drop-down list. If the file is not loaded yet, it will be shown in red. The first three files are Gene Ontology files. You can load other ontologies, but you need your own annotation file to annotate networks.
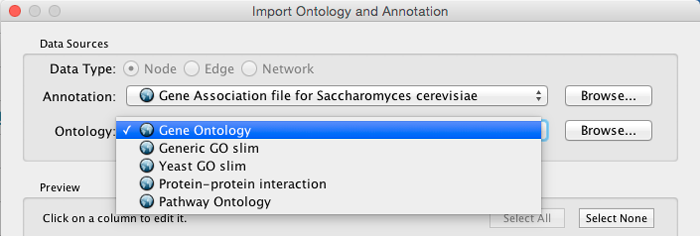
Once you click the Import button, Cytoscape will start loading OBO and Gene Association files from the remote sources. If you choose GO Full it may take a while since it is a large data file.
- When Cytoscape finishes importing files, the import window will be automatically closed. All columns mapped by this function have the prefix "annotation" and look like this: annotation.[column_name].
Note: Cytoscape supports both OBO formats: version 1.0 and 1.2.
Linkout
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
The external links are specified in a linkout.props file which is internal to Cytoscape. The defaults include a number of links such as Entrez, SGD, iHOP, and Google, as well as a number of species-specific links. In addition to the default links, users can customize the External Links menu and add (or remove) links by editing the linkout properties (found under Edit → Preferences → Properties...).
External links are listed as ‘key’-‘value’ pairs in the linkout.props file where key specifies the name of the link and value is the search URL. The LinkOut menus are organized in a hierarchical structure that is specified in the key. Linkout key terms specific for nodes start with the keyword nodelinkouturl, for edges this is edgelinkouturl.
For example, the following entry:
nodelinkouturl.Model Organism DB.SGD (yeast)=http://www.yeastgenome.org/cgi-bin/locus.fpl?locus=%ID%
places the SGD link under the Model Organism DB submenu. This link will appear in Cytoscape as:
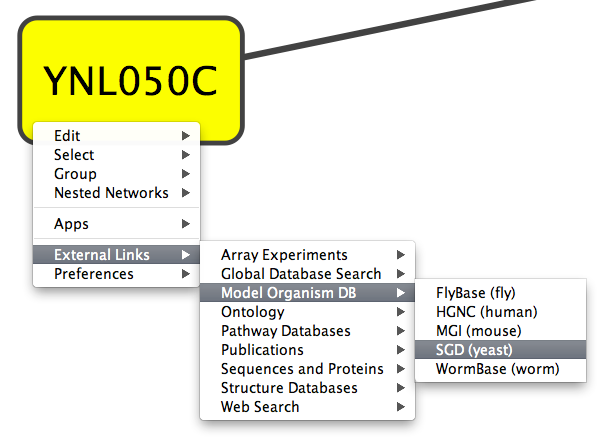
In a similar fashion one can add new submenus.
The %ID% string in the URL is a place-holder for the node label. When the popup menu is generated this marker is substituted with the node label. In the above example, the generated SGD link for the YNL050C protein is:
http://www.yeastgenome.org/cgi-bin/locus.fpl?locus=YNL050C
If you want to query based on a different column, you need to specify a different node label using Styles.
For edges the mechanism is much the same; however here the placeholders %ID1% and %ID2% reflect the source and target node label respectively.
Currently there is no mechanism to check whether the constructed URL query is correct and if the node label is meaningful. Similarly, there is no ID mapping between various identifiers. For example, a link to NCBI Entrez from a network that uses Ensembl gene identifiers as node labels will produce a link to Entrez using the Ensembl ID, which results in an incorrect link. It is the user's responsibility to ensure that the node label that is used as the search term in the URL link will result in a meaningful link.
Adding and Removing Links
The default links are defined in a linkout.props file contained inside the Linkout JAR bundle under the framework/system/org/cytoscape/linkout-impl subdirectory of the Cytoscape installation. These links are normal Java properties and can be edited by going to Edit → Preferences → Properties... and selecting linkout from the box (shown below). Linkouts can be modified, added or removed using this dialog; however, note that the modifications would not be stored in the file. To change a URL permanently, you would need to edit the linkout.props file directly.
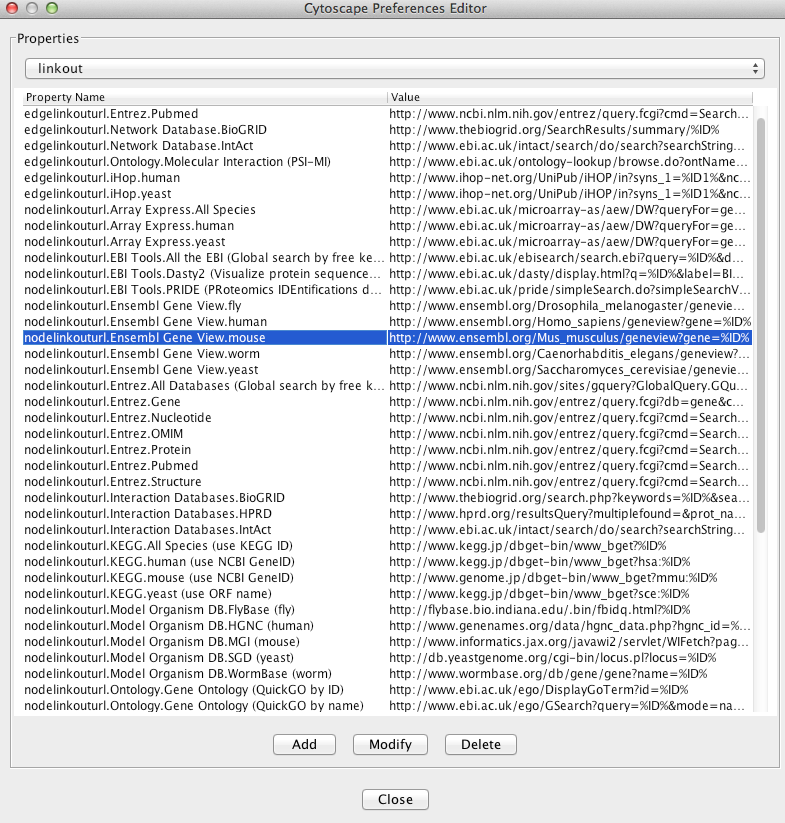
In addition, new links can be defined when starting Cytoscape from command line by specifying individual properties. The formatting of the command is cytoscape.sh -P [context_menu_definition]=[link] . context_menu_definition specifies the context menu for showing the linkout menu item. The structure of this definition is "." separated and the first item needs to be either nodelinkouturl or edgelinkouturl. The former will add the linkout item as a node context menu and the latter will add it as an edge context menu. The rest of the definition would define the hierarchy of the menu.
For instance this command:
cytoscape.sh -P nodelinkouturl.yeast.SGD=http://db.yeastgenome.org/cgi-bin/locus.pl?locus\=%ID%
will add this menu item:
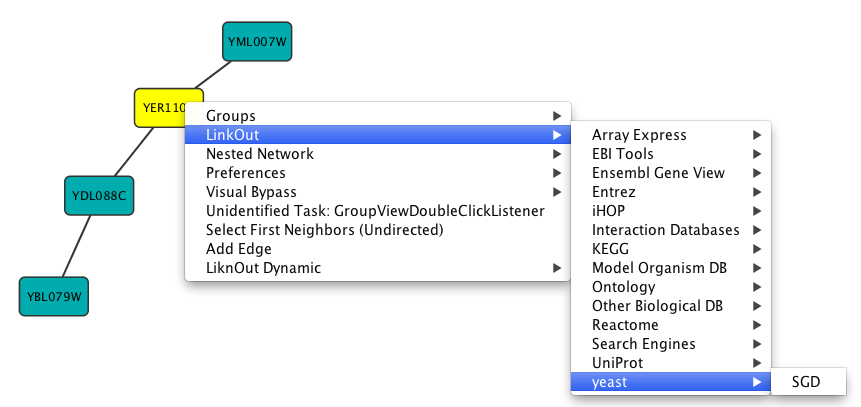
To remove a link from the menu, simply delete the property using Edit → Preferences → Properties... and selecting commandline. Linkouts added in the command line will be available for the running instance of Cytoscape.
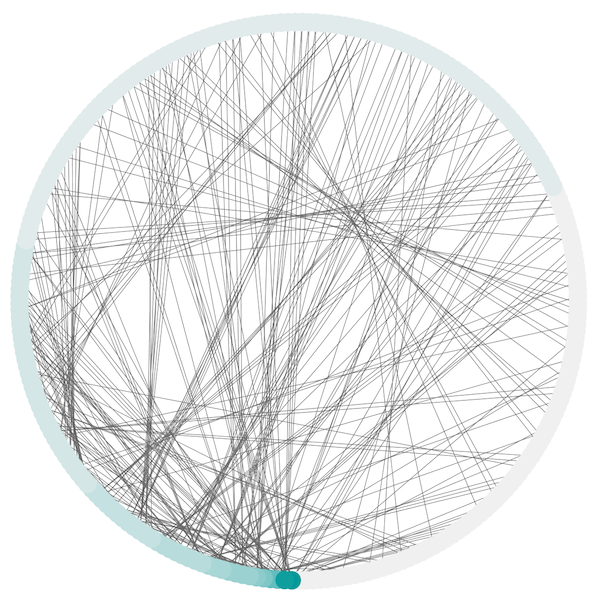
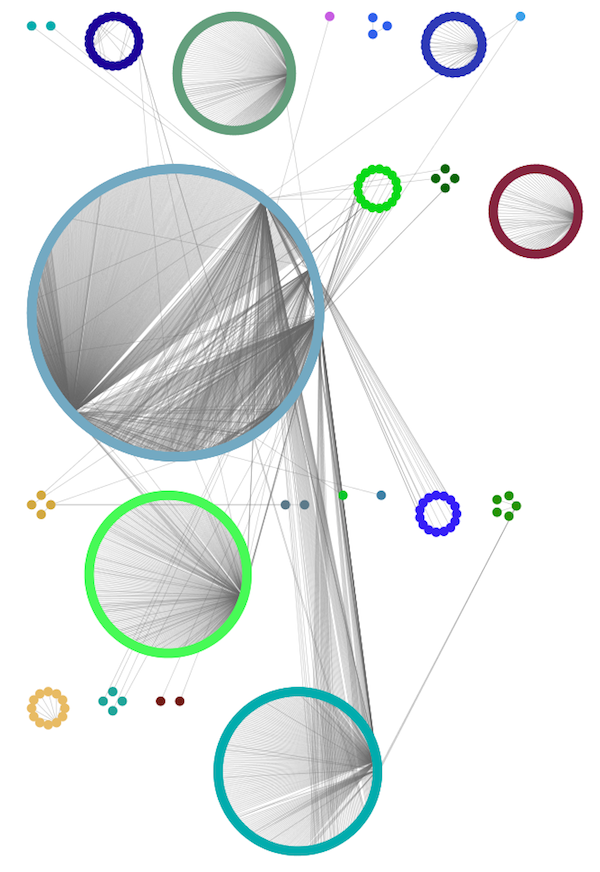
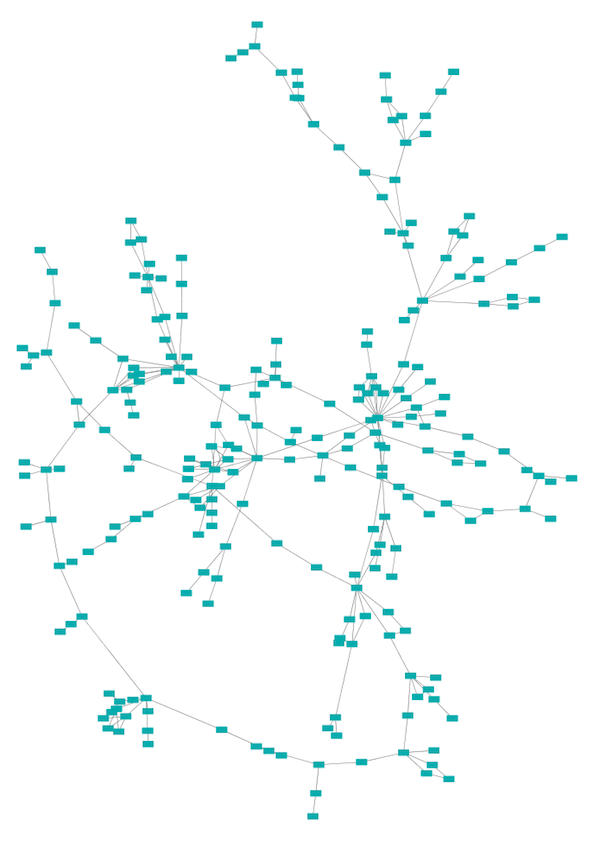
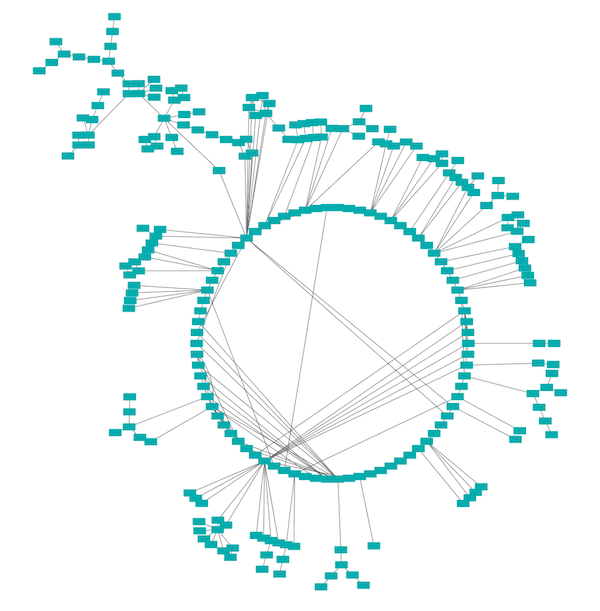
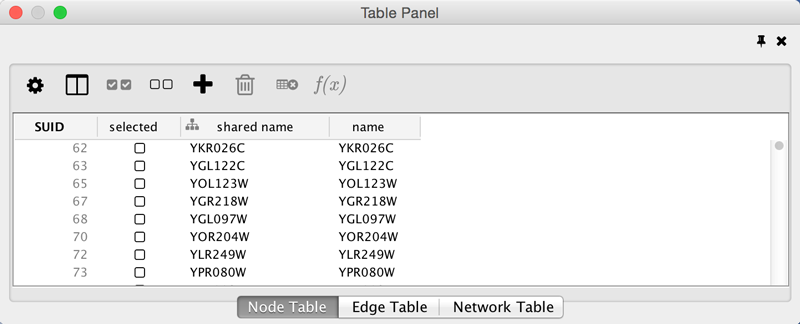
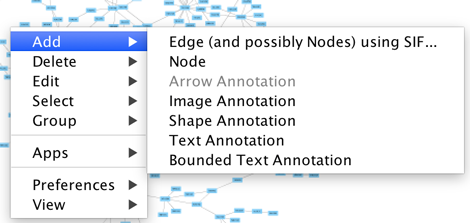
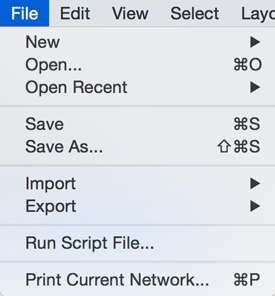
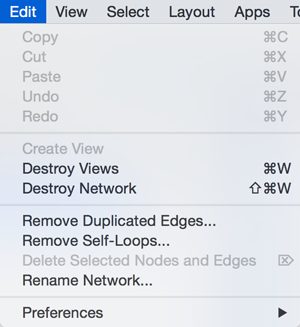
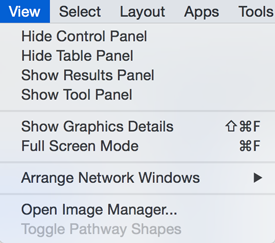
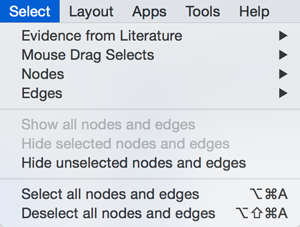
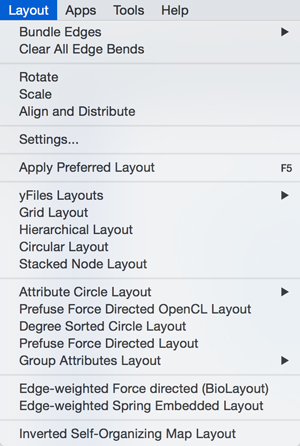

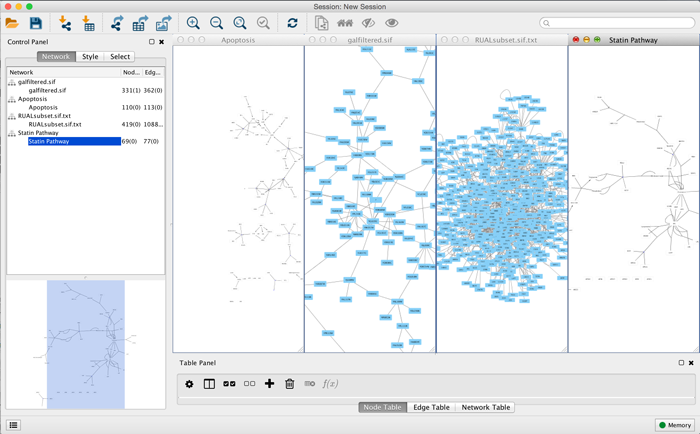
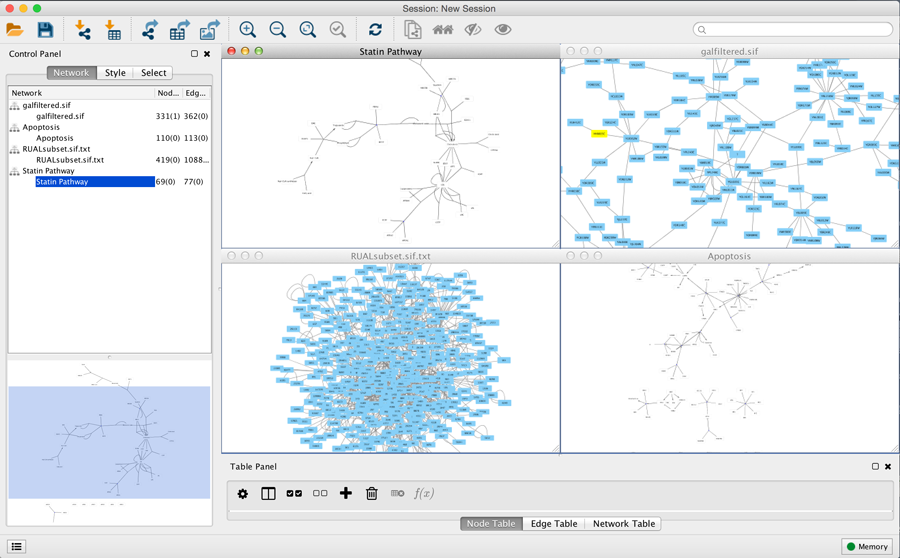
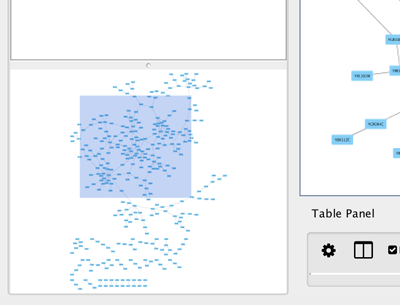
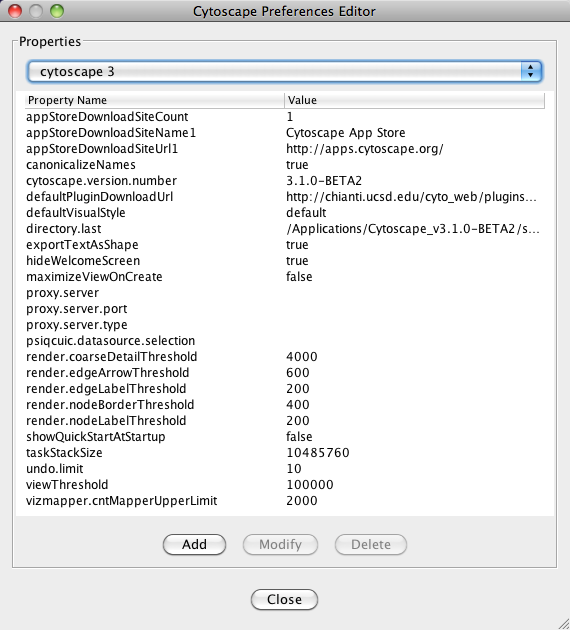
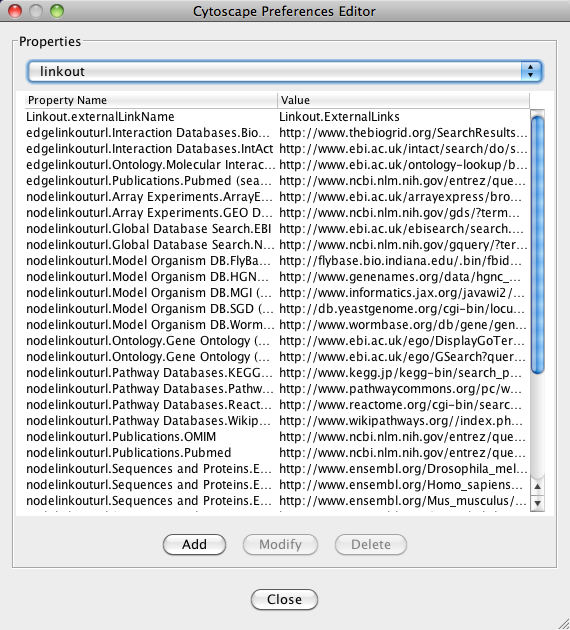
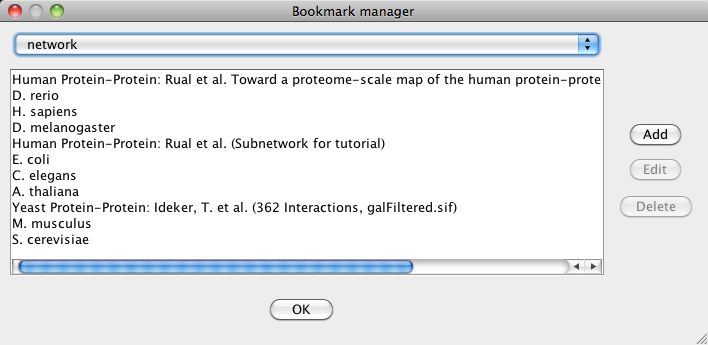
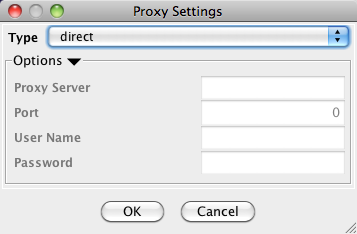
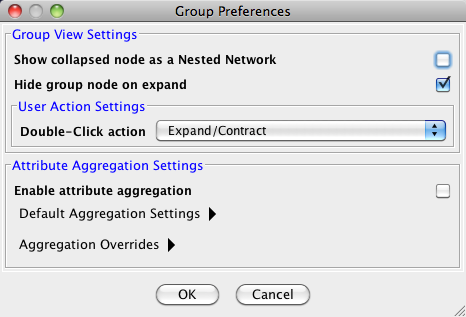
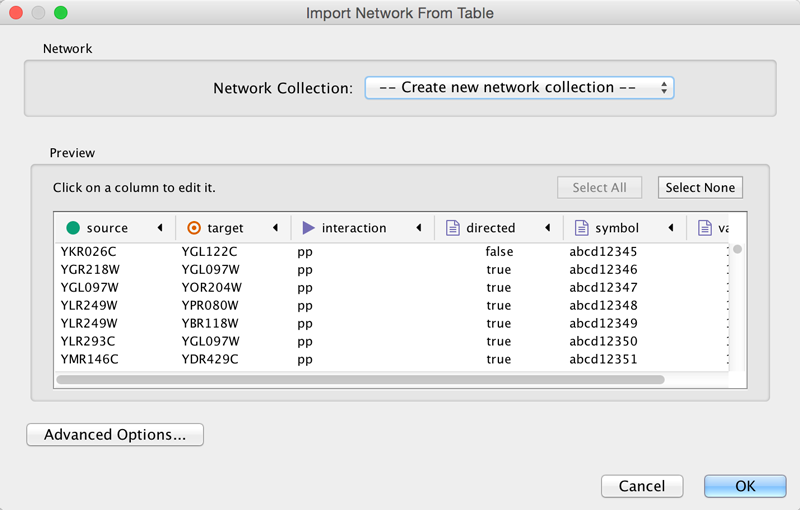
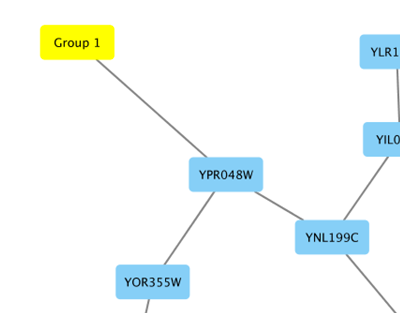
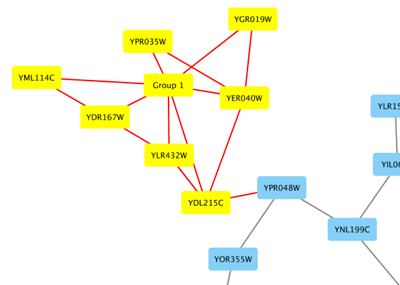
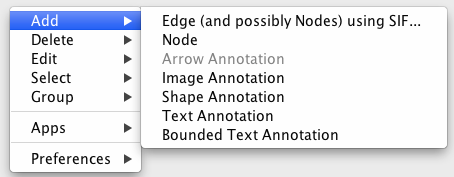






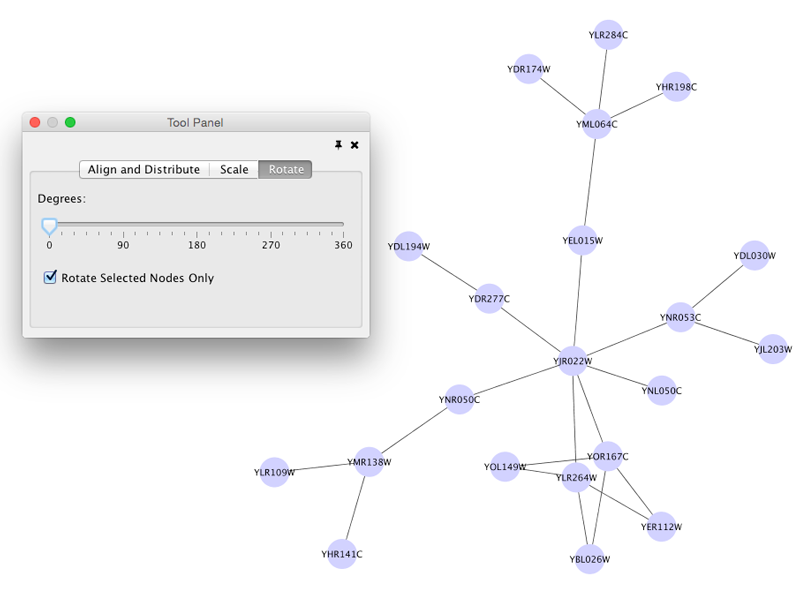
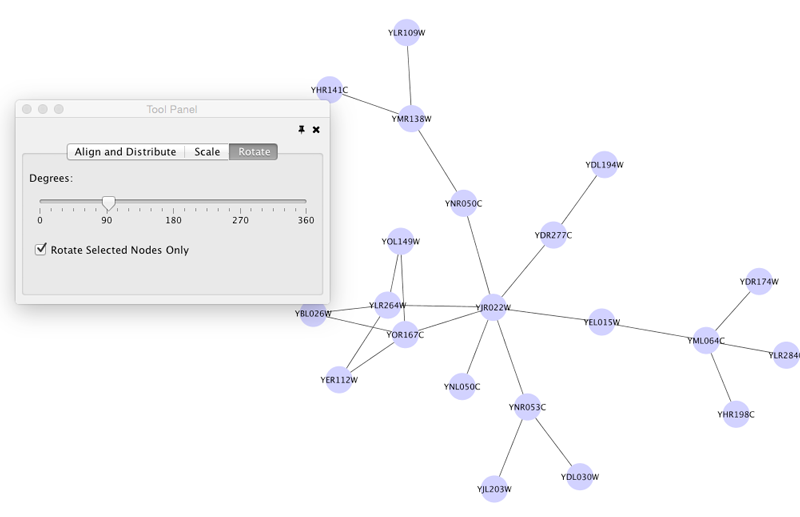
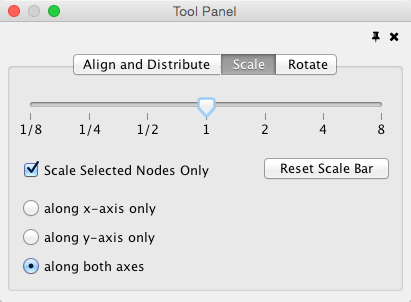
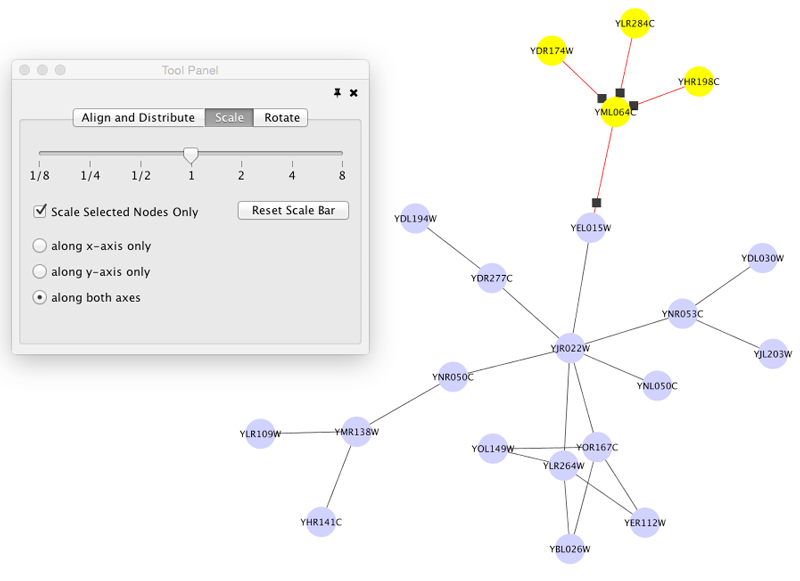
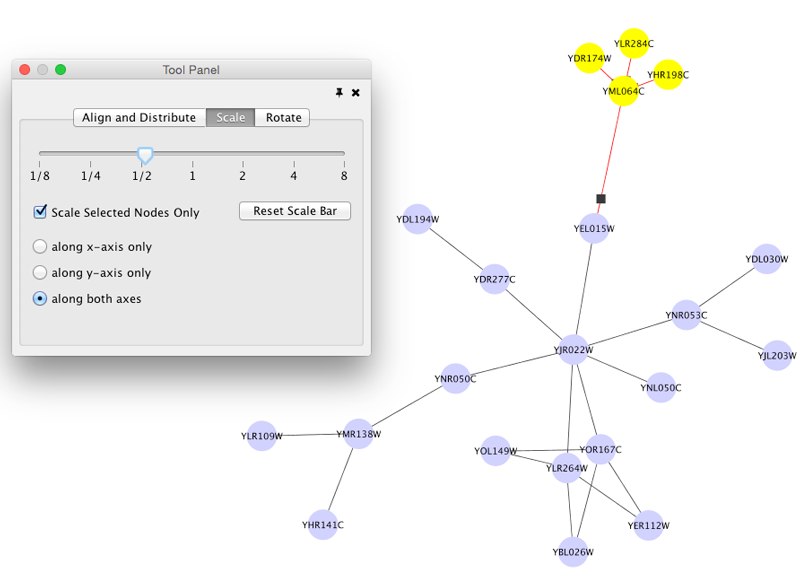
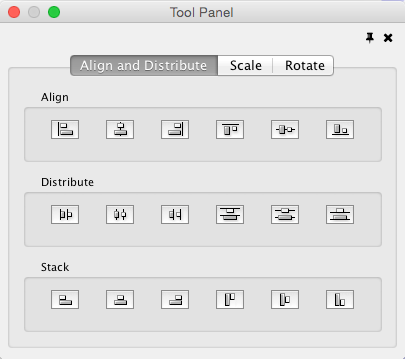

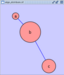





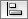





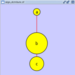



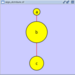

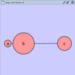
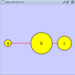
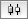
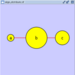

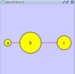

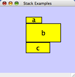

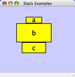

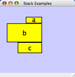

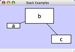
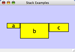

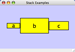

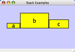


The Update Organism Preset Networks option updates the preset networks available on the Welcome screen to the latest version.