Cytoscape 3.0 Use Cases
Please write the use cases in terms of Cytoscape, meaning talk about node, edges, and attributes rather than talking about proteins and interactions. Instead, use specific biological discussions to illustrate instances of particular use cases. Try to be as general as possible. Try to see how your use case might match another use case that already exists.
Use Case: Biochemical Reactions (HyperEdges)
A Cytoscape Edge typically connects two Nodes, which we commonly refer to as Source and Target Nodes. While this is sufficient for most cases, there are cases where a user may want to have Edges that connect more than two Nodes. One example of this is a Biochemical Reaction, where an Edge represents a reaction that may have multiple substrates, products, and mediators. An example of this is shown in the figure below,
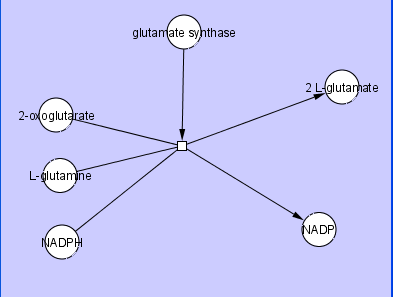
where L-glutamine and 2-oxoglutarate are substrates for the reaction, the catalyst glutamate substrate is a mediator for the reaction, 2 L-glutamate is a product of the reaction, the co-factor NADPH is a substrate for the reaction, and the co-factor NADP is a product of the reaction.
There may also be cases in which a user may want to connect one edge with another edge, for example to represent the activity of a molecular species that modulates the action of a catalyst. This is illustrated in the figure below
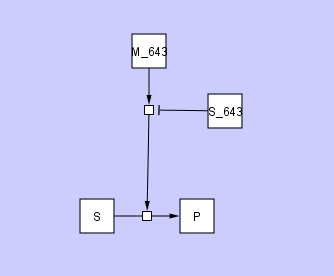
where molecular species S_643 inhibits the action of catalyst M_643.
If you look closely at these two figures, you will see some small squares at the intersections of the edges. We refer to this small square as a ConnectorNode. We also refer to the collection of edges and ConnectorNode as a HyperEdge.
The following two figures show examples of biochemical reactions -- Krebs Cycle and Glycolysis Reaction.
The first figure below illustrates Glycolosis Reaction. Note the use of shared edges and multiple connections to the same Node within a HyperEdge.
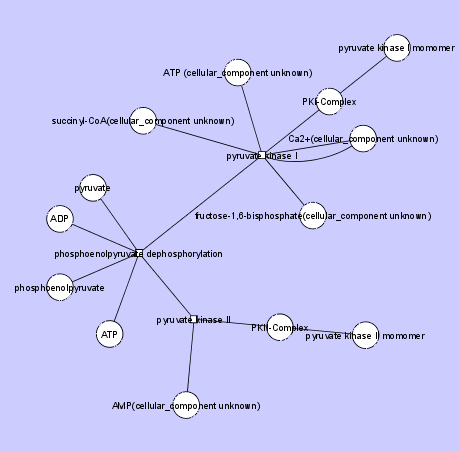
The second example below uses a Circle Graph Layout to illustrate the Krebs Cycle.
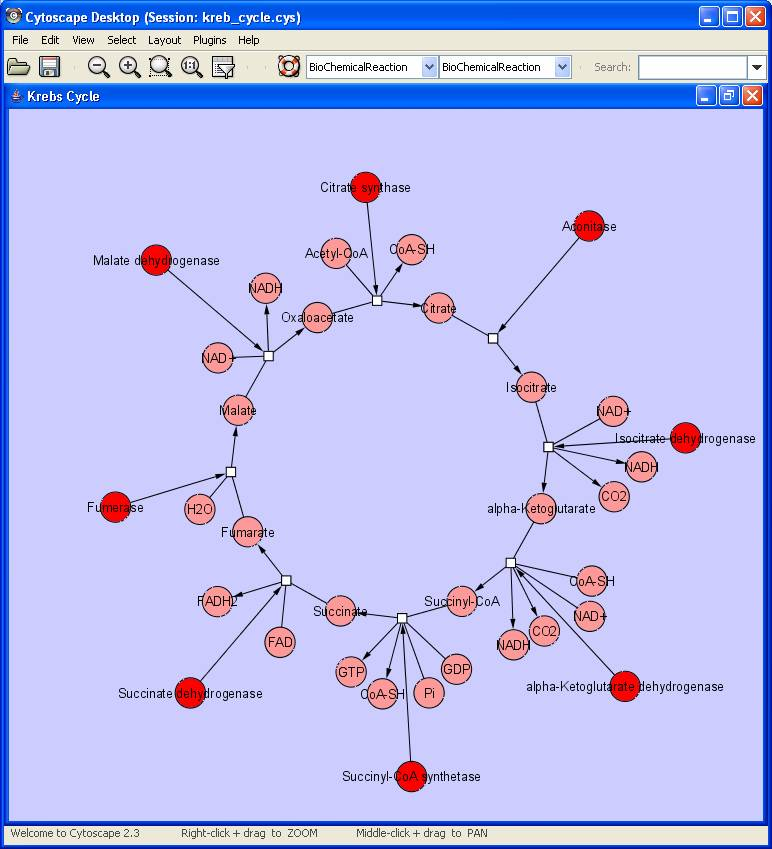
A detailed discussion of HyperEdge design issues can be found on the wiki page at Cytoscape_3.0/HyperEdges.
Use Case: Scientific Illustration
Cytoscape is used by researchers who need to publish and present their work. Often a user will want to include figures of Cytoscape networks in their publications and/or presentations. This requires presenting the Cytoscape network in a way that is intuitive to biologists. But Cytoscape networks are designed for computation, not presentation. The figure below shows a segment of a Cytoscape network for Wnt signalling, imported originally from BioPAX.
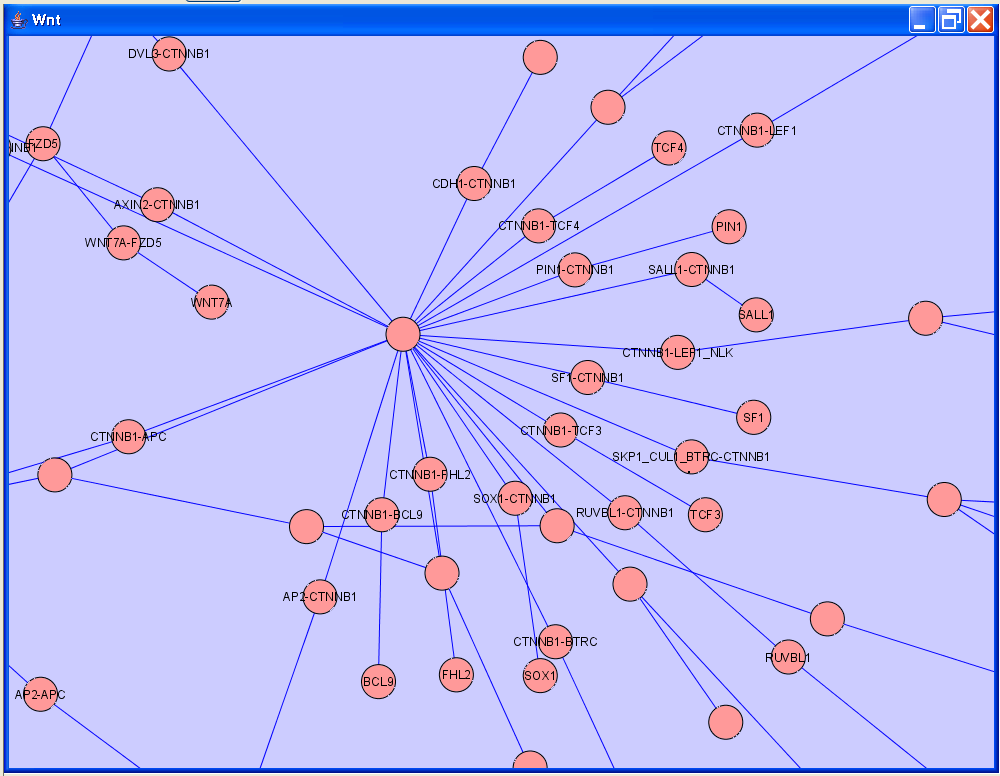
Contrast this with a manually illustrate Wnt signalling pathway, as shown in the figure below, derived from Wikipathways (http://www.wikipathways.org).
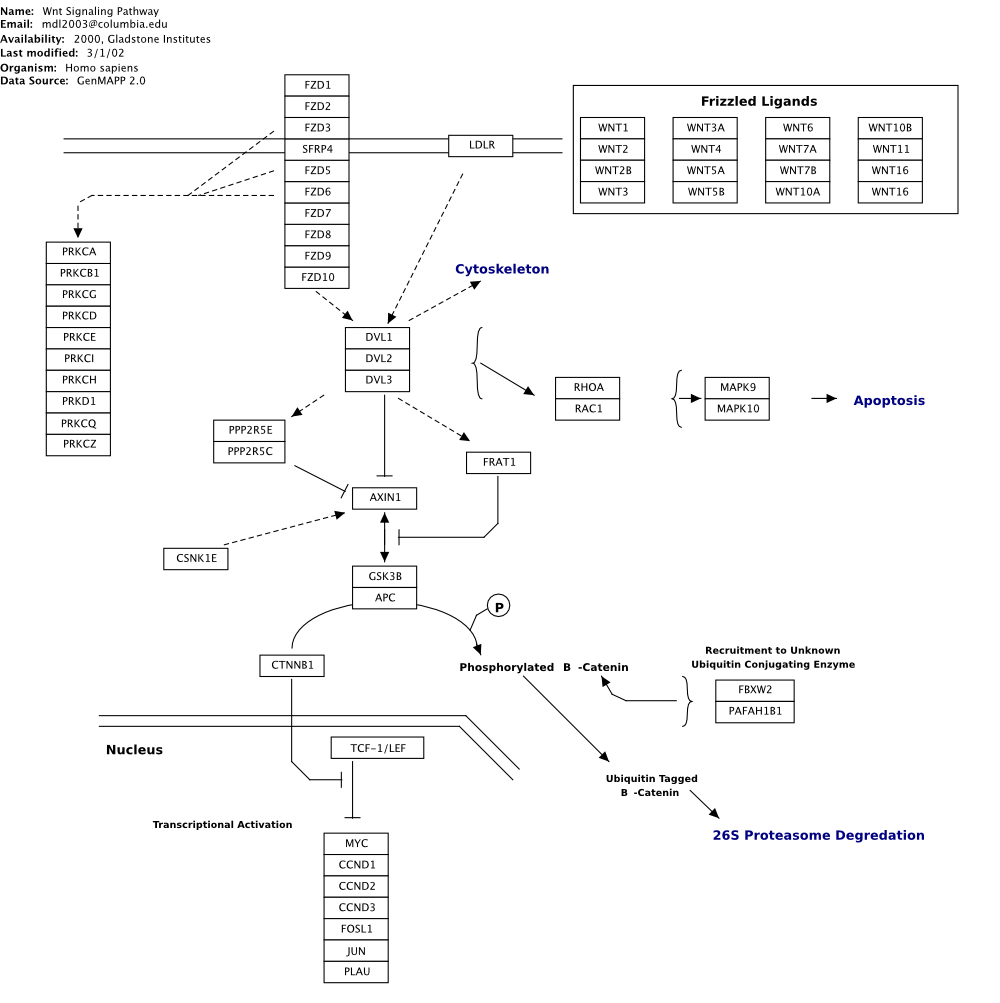
Note the use of annotational elements such as brackets, free text, and free-form sketched objects, such as the nuclear membrane shown in bottom left of the figure.
Cytoscape 3.0 should support both styles of network views. To do this, there needs to be support for arbitrary graphical and textual annotation. Users should be able to define new shapes. We should define a set of constraints such that user-defined shapes that hold to those constraints can be supported by the underlying Cytoscape renderer. Users should also have control over how different shapes will overlap. To this end, we have begun to integrate the construct of multiple Layers into the Cytoscape canvas class.
The figure below shows an initial implementation of an interface between Wikipathways and Cytoscape, written by Thomas Kelder at University of Maastricht, which imports a network written in the GPML markup format that underlies Wikipathways. Note the use of custom shapes, such as brackets, and free form text.
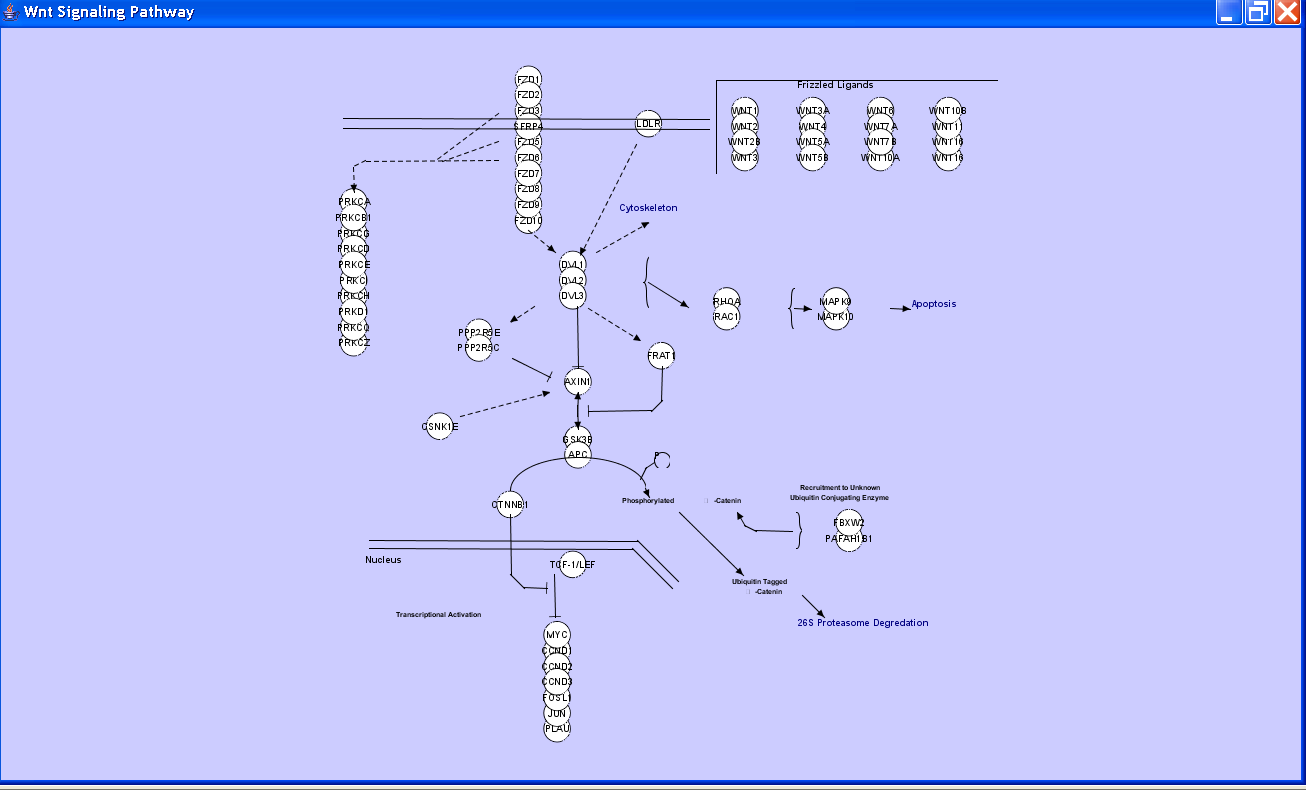
Note the use of custom shapes, such as brackets, and free form text. We should discuss the implications of implementing such constracts more directly in Cytoscape, from the perspectives of performance, expressiveness, and usability.