<<TableOfContents: execution failed [Argument "maxdepth" must be an integer value, not "[2]"] (see also the log)>>
Summary
A user wants to describe the possibility that a protein molecule (modified or unmodified) binds with an identical molecule of the same protein to form a dimer.
Step-by-Step User Action
- User specifies a modified or unmodified protein that can dimerize
- User specifies that a dimer is formed
- User optionally specifies the evidence for this reaction
Visual Aides
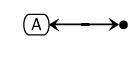
Here is an example of the Kohn notation of a homodimer follows:
Requirements for Cytoscape
TBD
Importance
Homodimerization is a very common biological process.
Variations
If the homodimer is used in a subsequent reaction, then a dot is placed in the middle. The dot in this case, represents the complex A:A.
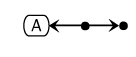
In addition to homodimers of unmodified proteins, modified proteins can also be represented forming homodimers:
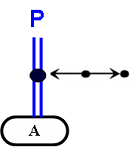
The color in this example is convention, and is not an essential element in the notation.
Other Examples
Comments
Shared ../MimEditorUseCaseComments
A Homodimer could be represented in BioPAX as a Complex object (e.g. Complex object with Components = A,A, where A is a physicalEntity)
AllanKuchinsky - 2006-12-11 03:40:57
At the model level, do we represent this as an edge whose source and target point to same node? If so, then this can be supported under current model and the only modifications would be those needed at the view level. Currently Cytoscape would render this as a self-connecting node, so modifications would be needed to use the dot notation.
In the first variation, are we connecting to the homodimer itself, rather than one of its elements? If so, then we should use a Group to represent this, much as we'd use a Group to represent a complex.
In the second variation, is the dot at right semantically very different from the dot in the middle of the modification reaction, the former representing a species and the latter representing a binding reaction?
MiritAladjem - 2006-12-13 12:26:14
Thank you Allan for the comments.
Regarding the first question, it is possible to represent homodimer formation as an edge whose source and target point to the same node.
Regarding the second question, in most cases the connections (e.g. non-covalent binding) will be to the homodimer itself. A group representation of a complex will be appropriate. There will be however some cases in which the connections will be to one of the components of the dimer (e.g. a covalent modification on one of the two monomers that interact to create the homodimer), so this option needs to be available.
Regarding the third questiohn, yes the dot at right is very different from the dot in the middle. The dot on the right represents a second copy of the molecular species on the left (here designated as A) whereas the dot in the middle represent the complex (in this example, a complex of two copies of A).
AllanKuchinsky - 2007-05-25 09:39:50
Regarding the second variation, does the dot at right represent the molecular species in its unmodified or modified state?