Use Case Name : ../ProteinBindingSites |
For Feature : MIMEditor |
Editors: DavidKane |
<<TableOfContents: execution failed [Argument "maxdepth" must be an integer value, not "[2]"] (see also the log)>>
Summary
A user wants to describe interactions of a protein, binding site by binding site.
Step-by-Step User Action
- User specifies a protein to be described in more detail
- User specifies the number of binding sites to be included
- User specifies the name of each binding site
- User then uses each binding site in subsequent reactions
Visual Aides
Binding sites are indicated with the binding site label sticking out perpendicularly from the protein. An example follows:

A binding site may also be used in combination with the specification of ../ProteinDomains:
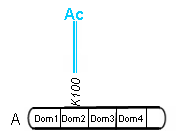
Requirements for Cytoscape
TBD
Importance
This is a moderately important feature of the notation.
Other Examples
Comments
Shared ../MimEditorUseCaseComments
BioPax representation TBD
GaryBader 2006-12-26 12:28:20
Similar to sub-gene use case for Group API
AllanKuchinsky 2007-01-22 05:11:24
case of binding site label sticking out perpendicularly from the protein can be handled with Custom Node Graphic