Use Case Name : ../DnaSite |
For Feature : MIMEditor |
Editors: DavidKane |
<<TableOfContents: execution failed [Argument "maxdepth" must be an integer value, not "[2]"] (see also the log)>>
Summary
A user wants to describe a DNA site that is involved in a pathway
Step-by-Step User Action
- User specifies that there is a DNA site
- User labels the DNA site
Visual Aides
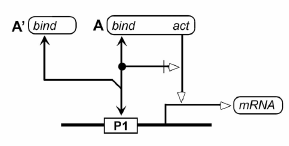
The notation for a DNA site is the following:

An example using this notation follows:
Requirements for Cytoscape
TBD
Importance
TBD
Other Examples
Comments
Shared ../MimEditorUseCaseComments
BioPax representation TBD
GaryBader - 2006-12-26 12:12:38
This is related to the general problem of representing sequence features in the context of a network. BioPAX Level 2 has basic support for describing sequence features (taken from PSI-MI), however Level 3 will add support for stating which sequence features are bound to others (related to describing topology of a complex).
GaryBader - 2006-12-26 12:13:16
Also, this is related to the sub-gene use case by the GenMAPP group at http://cytoscape.org/cgi-bin/moin.cgi/groupAPI/UseCase_3A
AllanKuchinsky - 2007-01-19 03:37:08
What is the formal relationship between A and A'?
MiritAladjem - 2007-02-01 08:58:10
In answer to Alan's comment: A' is a variant of A that has the binding site but not the acting site. There are real examples of such molecules (e.g. splicing variants). In the depicted case, A or A' can both bind DNA but they compete on the same site so just one of those two molecules can bind to the site at a specific time.