Use Case Name : Protein Complexes |
For Feature : Groups |
Editors: Alex Pico |
<<TableOfContents: execution failed [Argument "maxdepth" must be an integer value, not "[2]"] (see also the log)>>
Summary
We would like to be able to represent two or more proteins as macromolecular complexes in pathways. The representation of complexes will help simplify the visualization of complicated biological systems. Furthermore, complexes themselves have properties and interactions that we'd like to support and store.
Step-by-Step User Action
Creating a New Complex
- Select two or more nodes
- Choose "Form complex" from a context menu, main menu or toolbar
- Automatically view collapsed view of complex with default label (editable)
- Be able to expand complex as vertically stacked set of nodes by a very simple mechanism (e.g., click on a plus/minus icon)
- perhaps other restricted views of the children will be allowed:
- horizontal stack
- block (e.g., 2x2, 3x4)
- overlapping "clump" of nodes packed into a some defined circular area
- new network
- perhaps other restricted views of the children will be allowed:
- Be able to destroy complex
- Be able to expand-all or collapse-all complexes in a given network
Loading a Network with a Complex
- Same as 3-6 above
Visual Aides
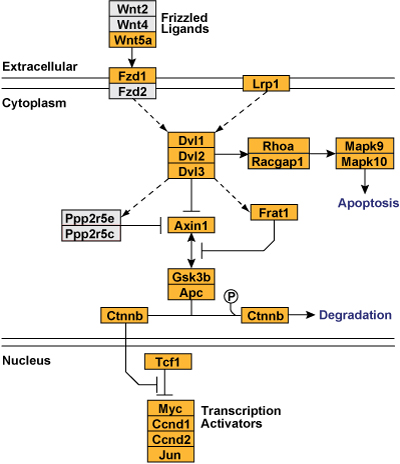
In this GenMAPP pathway, stacked gene objects help indicate complex and paralog groups. For example, Rhoa + Racgap1 and Gsk3b + Apc
Requirements for Cytoscape
- The Group API can handle this one in terms of create, expand, collapse and destroy
- Need to implement a simple GUI that is specific for complexes, i.e., uses semantics relating to protein complexes
- Need to add simple layout/alignment algorithms for stacking children vertically in expanded view
- Need to support the storage of complexes in a pathway file format. xGMML and GenMAPP's GPML?
Importance
This is a very important use case, critical to almost every pathway. The typical user would expect quick, intuitive control over this feature, mainly for simplifying (or digging deeper into) a given pathway representation.
Other Examples
- Simple, intuitive examples of expand and collapse can be found in directory tools/explorers that let you click on an icon (e.g., a triangle or plus/minus) to expand and collaspe the view.
- Illustrator and other such programs have a grouping function that mimics the restriction on children once a group is formed. You cannot move the children relative to eachother, for example. If you want to treat them independently, you have to ungroup.
Comments
AllanKuchinsky - 2006-11-27 07:16:15
This suggests a sub-project to define a set of "visual formalisms" for the view aspect of Groups. Visual formalisms are "diagrammatic notations with well-defined semantics for expressing relations. They ae based on simple visual notations such as tables, graphs, plots, panels, and maps. Versions of such visual notations that define a precise semantics become truly formal". This is described in Nardi and Zarmer, 1990, which is attached nardiZarmer.pdf
AllanKuchinsky - 2006-11-27 07:33:55
The idea would be to build a small set of visual formalisms, e.g. vertical/horizontal stacks, tables, that could be readily specialized to views for complexes, paralogs, protein domains, etc.