Cytoscape is a Java application verified to run on Linux, Windows, and Mac OS X. Although not officially supported, other UNIX platforms such as Solaris or FreeBSD may run Cytoscape if Java version 5 or later is available for the platform.
System requirements
The system requirements for Cytoscape depend on the size of the networks the user wants to load, view and manipulate.
|
Small Network Visualization |
Large Network Analysis/Visualization |
Processor |
1GHz |
As fast as possible |
Memory |
512MB |
2GB+ |
Graphics Card |
On board Video |
Highend Graphics Card |
Monitor |
XGA (1024X768) |
Wide or Dual Monitor |
Getting Started
Install Java
If not already installed on your computer, download and install Java SE 5 or 6. Cytoscape 2.5 will no longer run with Java version 1.4.x or lower. You must install Java SE 5 or 6!!!
These can be found at:
In general, Java SE 6 is faster than 5. If your machine is compatible with the 6 series, please try version 6.
Install Cytoscape
A quick note on upgrading
There should really be no issue in upgrading. If you have a previous installation you have two options: 1) Starting with a clean slate: For this you should delete your previous installation directory and the .cytoscape directory. 2) Just keep what you have and simply pick a distinct, new directory for installation. In the unlikely event that you should encounter any problem, delete the .props files in your .cytoscape directory. If that doesn't help try deleting the .cytoscape directory. This latter step will cause you to loose all your plugins that you have installed via the plugin manager, so only do that if you are having problems or if you don't mind reinstalling your plugins. The core plugins will not be affected by this step.
There are a number of options for downloading and installing Cytoscape. All options can be downloaded from the http://cytoscape.org website.
- Automatic installation packages exist for Windows, Mac OS X, and Linux platforms.
- You can install Cytoscape from a compressed archive distribution.
- You can build Cytoscape from the source code.
You can check out the latest and greatest software from our Subversion repository.
Cytoscape installations (regardless of platform) containing the following files and directories:
File |
Description |
cytoscape.jar |
Main Cytoscape application (Java archive) |
cytoscape.sh |
Script to run Cytoscape from command line (Linux, Mac OS X) |
cytoscape.bat |
Script to run Cytoscape (Windows) |
LICENSE.txt/html |
Cytoscape GNU LGPL License |
lib/ |
library jar files needed to run Cytoscape. |
docs/ |
Manuals in different formats. What you are reading now. |
licenses/ |
Licence files for the various libraries distributed with Cytoscape. |
plugins/ |
Directory containing cytoscape plugins, in .jar format. |
sampleData/ |
|
|
galFiltered.gml -- Sample molecular interaction network file * |
|
galFiltered.sif -- Identical network in Simple Interaction Format * |
|
galExpData.pvals -- Sample gene expression matrix file * |
|
galFiltered.nodeAttrTable.xls -- Sample node attribute file in Microsoft Excel format |
|
galFiltered.cys -- Sample session file created from datasets above plus annotations from several databases * |
|
BINDyeast.sif -- Network of all yeast protein-protein interactions in the BIND database as of Dec, 2006 ** |
|
BINDhuman.sif -- Network of all human protein-protein interactions in the BIND database as of Dec, 2006 ** |
|
yeastHighQuality.sif -- Sample molecular interaction network file *** |
|
interactome_merged.networkTable.gz -- Human interactome network file in tab-delimited format **** |
|
sampleStyles.props -- Additional sample Visual Styles |
* From Ideker et al., Science 292:929 (2001)
** Obtained from data hosted at http://www.blueprint.org/bind/bind_downloads.html
** From von Mering et al., Nature, 417:399 (2002) and Lee et al, Science 298:799 (2002)
**** Created from Cytoscape tutorial web page. Original data sets are available at: http://www.cytoscape.org/cgi-bin/moin.cgi/Data_Sets/ from "A merged human interactome" by Andrew Garrow, Yeyejide Adeleye and Guy Warner (Unilever, Safety and Environmental Assurance Center).
Launch the application
Double-click on the icon created by the installer or by running cytoscape.sh from the command line (Linux or Mac OS X) or double-clicking cytoscape.bat (Windows). Alternatively, you can pass the .jar file to Java directly using the command java -Xmx512M -jar cytoscape.jar -p plugins. The -Xmx512M flag tells java to allocate more memory for Cytoscape and the -p plugins option tells cytoscape to load all of the plugins in the plugins directory. Loading the plugins is important because many key features like layouts, filters and the attribute browser are included with Cytoscape as plugins in the plugins directory. See the Command Line chapter for more detail. In Windows, it is also possible to directly double-click the .jar file to launch it. However, this does not allow specification of command-line arguments (such as the location of the plugin directory).
When you succeed in launching Cytoscape, a window will appear that looks like this (captured on Mac OS 10.4):
Note on Memory Consumption
For users interested in loading large networks, the amount of memory needed by Cytoscape will increase. Memory usage depends on both number of network objects (nodes+edges) and the number of attributes. Here are some rough suggestions for memory allocation:
Suggested Memory Size Without View
Number of Objects (nodes + edges) |
Suggested Memory Size |
0 - 70,000 |
512M (default) |
70,000 - 150,000 |
800M |
Suggested Memory Size With View
Number of Objects (nodes + edges) |
Suggested Memory Size |
0 - 20,000 |
512M (default) |
20,000 - 70,000 |
800M |
70,000 - 150,000 |
1G |
Overall Memory Size for Cytoscape
To increase the maximum memory size for Cytoscape, you can specify it in a file, residing in the same directory as the Cytoscape executable, called Cytoscape.vmoptions with one option per line and lines separated by linefeeds. The last line must also be followed by a linefeed. The one exception to this rule is the MacOS platform if you are launching Cytoscape by clicking on the Cytoscape icon. (In that case you will have to edit the .../Cytoscape.app/Contents/Info.plist file instead!) For example, if you want to assign 1GB of memory, create the Cytoscape.vmoptions file containing the single line (Do not forget the linefeed at the end of the line!):
-Xmx1GB
Stack Size
There is one more option related to memory allocation. Some of the functions in Cytoscape use larger stack space (a temporary memory for some operations, such as Layout). Since this value is set independently from the Xmx value above, sometimes Layout algorithms fails due to the out of memory error. To avoid this, you can set larger heap size by using the -Xss option. If layout fails for large networks, please try the following:
-Xmx1GB -Xss10M
The option -Xss10M means set the heap size to 10MB. In many cases, this solves out of memory error for Layouts.
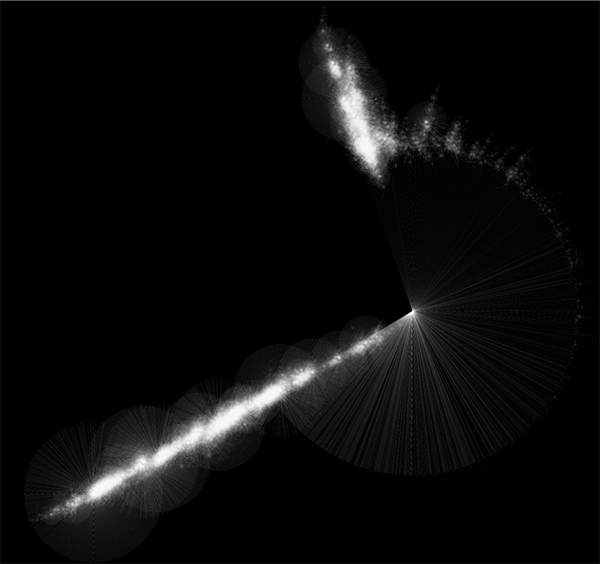
Randomly generated scale-free network with 500K nodes and 500k edges: If memory parameters are set properly, you can visualize huge networks. In this example, about 5GB of memory is used by Cytoscape. Stack size is set to 10MB. To use large memory space (4GB+) you need 64-bit version operating system AND 64-bit version Java SE 5 or 6.
Note: Some of the web service clients are multi-thread programs and each thread uses the memory size specified by -Xss option. If web service clients fails due to the out of memory error, please reduce the stack size and try again.
For more details, see How_to_increase_memory_for_Cytoscape.
Note on Directory Location
For the application to work properly, all files should be left in the directory in which they were unpacked. The core Cytoscape application assumes this directory structure when looking for the various libraries needed to run the application. If you are adventurous, you can get creative with the $CLASSPATH and/or the cytoscape.jar manifest file and run Cytoscape from any location you want.