Community Structure Layout
Gang Su
Bioinformatics Program, University of Michigan, Ann Arbor
Biological Use Case: This plugin is aimed to find community structures in large biological networks, which may infer protein complexes and biological pathways. We also utilize the community structure information to improve visualization of large interaction networks.
Recipe
Cytoscape version: Version number (2.6.1)
Plugins to Load: GLay.jar , MetaNodePlugin2.jar
GUI steps:
Describe each step (story), the GUI action to take, and probable remarks
Story |
Action |
Remarks |
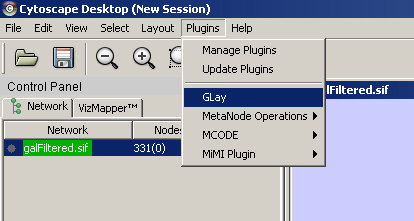
First of all, you need to load Glay. After you placed all the plugins jars in the folder, load Glay from the plugins tab. Also please load the preferred network. Here we used galFiltered.sif |
|
|
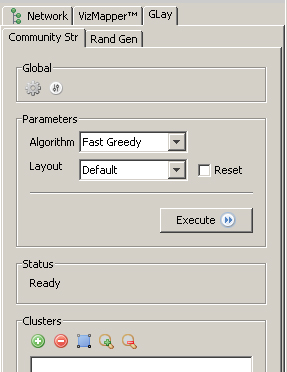
Then, you may tune the parameters. Right now we have two clustering algorithms and two layout algorithms. Just go with the default. |
|
|
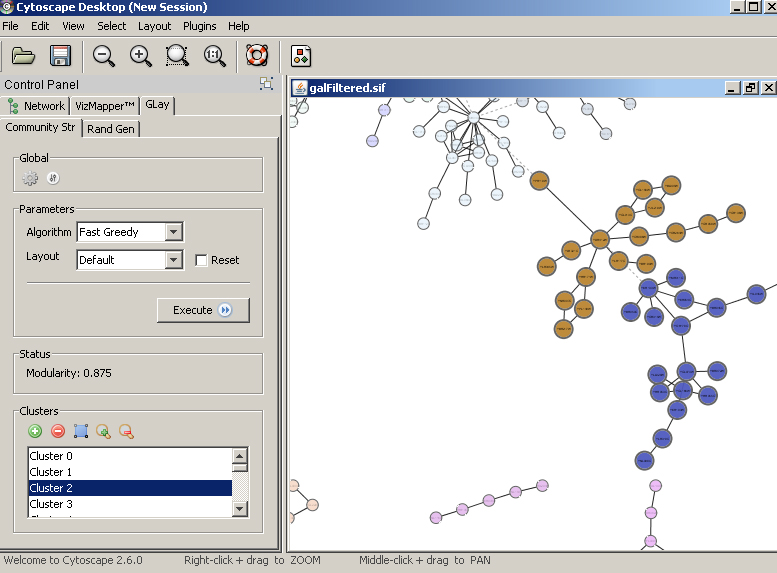
Next, you may interact with all the clusters from the cluster browsing panel; you may select multiple clusters from by holding the shift key. This will 'activate' clusters my changing visual styles. |
|
|
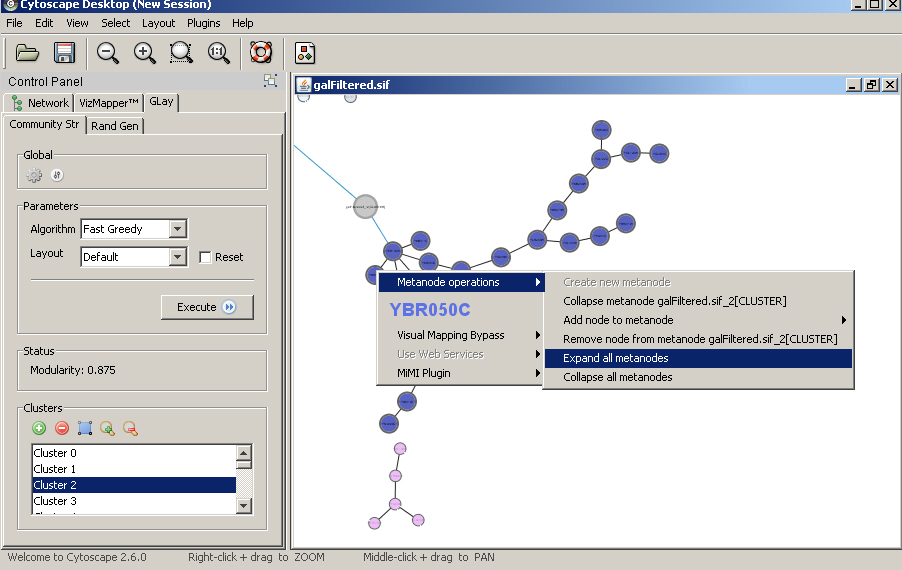
Clusters can be expanded or collapsed by right click on the nodes to bring up the contextual menu. |
|
|
Toolticons can be used to perform various tasks. |
|
|
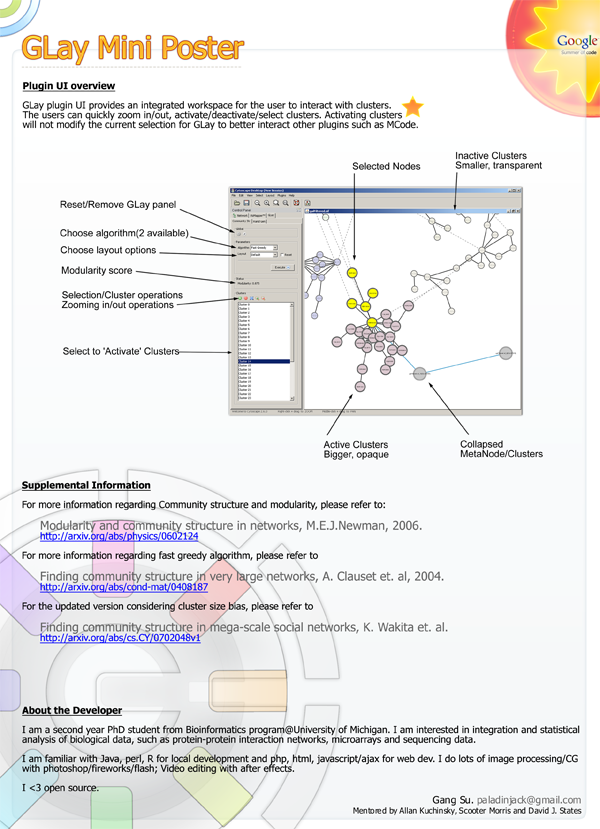
Here's a miniposter for FYI
And abstract FYI
Data / Session Files: Attach (preferably) a session file or else the data files used in the workflow
Presentation: Presentation.ppt
Webstart: Attach a webstart
Video: Video Tutorial: http://www.youtube.com/watch?v=W79Kb28348g, http://www.youtube.com/watch?v=OD3fUF62Gd8 ; High resolution video tutorial: GangScreen.wmv