|
← Revision 35 as of 2015-11-04 00:24:34
Size: 17576
Comment:
|
← Revision 36 as of 2016-04-26 22:15:14
Size: 61
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
|
Cytoscape can read network/pathway files written in the following formats: * Simple interaction file (SIF or .sif format) * Nested network format (NNF or .nnf format) * Graph Markup Language (GML or .gml format) * XGMML (extensible graph markup and modelling language). * SBML * BioPAX * PSI-MI Level 1 and 2.5 * GraphML * Delimited text * Excel Workbook (.xls, .xlsx) * [[http://cytoscape.github.io/cytoscape.js/#notation/elements-json|Cytoscape.js JSON]] The SIF format specifies nodes and interactions only, while other formats store additional information about network layout and allow network data exchange with a variety of other network programs and data sources. Typically, SIF files are used to import interactions when building a network for the first time, since they are easy to create in a text editor or spreadsheet. Once the interactions have been loaded and network layout has been performed, the network may be saved to GML or XGMML format for interaction with other systems. All file types listed (except Excel) are text files and you can edit and view them in a regular text editor. == SIF Format == The simple interaction format is convenient for building a graph from a list of interactions. It also makes it easy to combine different interaction sets into a larger network, or add new interactions to an existing data set. The main disadvantage is that this format does not include any layout information, forcing Cytoscape to re-compute a new layout of the network each time it is loaded. Lines in the SIF file specify a source node, a relationship type (or edge type), and one or more target nodes: {{{ nodeA <relationship type> nodeB nodeC <relationship type> nodeA nodeD <relationship type> nodeE nodeF nodeB nodeG ... nodeY <relationship type> nodeZ }}} A more specific example is: {{{ node1 typeA node2 node2 typeB node3 node4 node5 node0 }}} The first line identifies two nodes, called node1 and node2, and a single relationship between node1 and node2 of type typeA. The second line specifies three new nodes, node3, node4, and node5; here "node2" refers to the same node as in the first line. The second line also specifies three relationships, all of type typeB and with node2 as the source, with node3, node4, and node5 as the targets. This second form is simply shorthand for specifying multiple relationships of the same type with the same source node. The third line indicates how to specify a node that has no relationships with other nodes. This form is not needed for nodes that do have relationships, since the specification of the relationship implicitly identifies the nodes as well. Duplicate entries are ignored. Multiple edges between the same nodes must have different edge types. For example, the following specifies two edges between the same pair of nodes, one of type xx and one of type yy: {{{ node1 xx node2 node1 xx node2 node1 yy node2 }}} Edges connecting a node to itself (self-edges) are also allowed: {{{ node1 xx node1 }}} Every node and edge in Cytoscape has a name column. For a network defined in SIF format, node names should be unique, as identically named nodes will be treated as identical nodes. The name of each node will be the name in this file by default (unless another string is mapped to display on the node using styles). This is discussed in the section on '''[[Cytoscape_3/UserManual/Styles|Styles]]'''. The name of each edge will be formed from the name of the source and target nodes plus the interaction type: for example, {{{sourceName (edgeType) targetName}}}. The tag <edgeType> can be any string. Whole words or concatenated words may be used to define types of relationships, e.g. geneFusion, cogInference, pullsDown, activates, degrades, inactivates, inhibits, phosphorylates, upRegulates, etc. Some common interaction types used in the Systems Biology community are as follows: {{{ pp .................. protein – protein interaction pd .................. protein -> DNA (e.g. transcription factor binding upstream of a regulating gene.) }}} Some less common interaction types used are: {{{ pr .................. protein -> reaction rc .................. reaction -> compound cr .................. compound -> reaction gl .................. genetic lethal relationship pm .................. protein-metabolite interaction mp .................. metabolite-protein interaction }}} === Delimiters === Whitespace (space or tab) is used to delimit the names in the simple interaction file format. However, in some cases spaces are desired in a node name or edge type. The standard is that, if the file contains any tab characters, then tabs are used to delimit the fields and spaces are considered part of the name. If the file contains no tabs, then any spaces are delimiters that separate names (and names cannot contain spaces). If your network unexpectedly contains no edges and node names that look like edge names, it probably means your file contains a stray tab that's fooling the parser. On the other hand, if your network has nodes whose names are half of a full name, then you probably meant to use tabs to separate node names with spaces. Networks in simple interactions format are often stored in files with a {{{.sif}}} extension, and Cytoscape recognizes this extension when browsing a directory for files of this type. <<Anchor(nnf)>> == NNF == The NNF format is a very simple format that unlike SIF allows the optional assignment of single nested network per node. No other node columns can be specified. There are only 2 possible line formats: * A node "node" contained in a "network:" {{{network node}}} * 2 nodes linked together contained in a network: {{{network node1 interaction node2}}} If a network name (first entry on a line) appeared previously as a node name (in columns 2 or 4), the network will be nested in the node with the same name. Also, if a name that has been previously defined as a network (by being listed in the first column), later appears as a node name (in columns 2 or 4), the previously defined network will be nested in the node with the same name. In summary: any time a name is used as both, a network name , and a node name, this implies that the network will be nested in the node of the same name. Additionally comments may be included on all lines. Comments start with a hash mark '#' and continue to the end of a line. Trailing comments (after data lines) and entirely blank lines anywhere are also permissible. Please '''note''' that if you load multiple NNF files in Cytoscape they will be treated like a single, long concatenated NNF file! If you need to embed spaces, tabs or backslashes in a name, you must escape it by preceding it with a backslash, so that, e.g. an embedded backslash becomes two backslashes, an embedded space a backslash followed by a space etc. === Examples === ==== Example 1 ==== {{attachment:NNFExample1_2.png}} {{{ Example_1 C Example_1 network1 network1 A pp B network1 B pp A Example_1 C pp B }}} ==== Example 2 ==== {{attachment:NNFExample2_2.png}} {{{ Example_2 M1 Example_2 M2 M1 A M2 B pp C Example_2 A pp B Example_2 M1 im M2 }}} ==== Example 3 ==== {{attachment:NNFExample3_2.png}} {{{ Example_3 M1 im M2 Example_3 M3 im M1 Example_3 M2 im M3 Example_3 C pp M3 Example_3 M2 pp C M1 A M2 A pp B M3 B pp C }}} /* ==== Example 4 ==== */ /* {{attachment:NNFExample4.png}} */ /* Example_4 M1*/ /* root M3*/ /* M1 A pp B */ /* M1 B pp A */ /* Example_4 C pp B */ /* M3 M2 */ /* M2 D */ /* M3 E pp F */ /* M3 D pp F */ /* M3 D pp E */ /* Example_4 D pp C */ /* Example_4 A pp M2*/ /* Example_4 B pp M3 */ /* Example_4 M2 pp B */ ==== Example 5 ==== {{attachment:NNFExample5_2.png}} {{{ Example_5 M4 M4 D M4 M3 M3 M2 pp C M2 M1 pp B M1 A M4 C pp D }}} == GML Format == In contrast to SIF, GML is a rich graph format language supported by many other network visualization packages. The GML file format specification is available at: http://www.infosun.fmi.uni-passau.de/Graphlet/GML/ It is generally not necessary to modify the content of a GML file directly. Once a network is built in SIF format and then laid out, the layout is preserved by saving to and loading from GML. Properties specified in a GML file will result in a new style named {{{Filename.style}}} when that GML file is loaded. == XGMML Format == XGMML is the XML evolution of GML and is based on the GML definition. In addition to network data, XGMML contains node/edge/network column data. The XGMML file format specification is available at: http://cgi5.cs.rpi.edu/research/groups/pb/punin/public_html/XGMML/ XGMML is now preferred to GML because it offers the flexibility associated with all XML document types. If you're unsure about which to use, choose XGMML. There is a java system property "cytoscape.xgmml.repair.bare.ampersands" that can be set to "true" if you have experience trouble reading older files. This should only be used when an XGMML file or session cannot be read due improperly encoded ampersands, as it slows down the reading process, but this is still preferable to attempting to fix such files using manual editing. == SBML (Systems Biology Markup Language) Format == The Systems Biology Markup Language (SBML) is an XML format to describe biochemical networks. SBML file format specification is available at: http://sbml.org/documents/ == BioPAX (Biological PAthways eXchange) Format == BioPAX is an OWL (Web Ontology Language) document designed to exchange biological pathways data. The complete set of documents for this format is available at: http://www.biopax.org/ == PSI-MI Format == The PSI-MI format is a data exchange format for protein-protein interactions. It is an XML format used to describe PPI and associated data. PSI-MI XML format specification is available at: http://psidev.sourceforge.net/mi/xml/doc/user/ == GraphML == GraphML is a comprehensive and easy-to-use file format for graphs. It is based on XML. The complete set of documents for this format is available at: http://graphml.graphdrawing.org/ == Delimited Text Table and Excel Workbook == Cytoscape has native support for Microsoft Excel files (.xls, .xlsx) and delimited text files. The tables in these files can have network data and edge columns. Users can specify columns containg source nodes, target nodes, interaction types, and edge columns during file import. Some of the other network analysis tools, such as igraph (http://cneurocvs.rmki.kfki.hu/igraph/), has feature to export graph as simple text files. Cytoscape can read these text files and build networks from them. For more detail, please read the Import Free-Format Tables section of the '''[[Cytoscape_3/UserManual#Creating_Networks|Creating Networks]]''' section. == Cytoscape.js JSON == From Cytoscape 3.1.0 on, Cytoscape supports [[http://cytoscape.github.io/cytoscape.js/|Cytoscape.js]] JSON files. You can use this feature to export your network visualizations to web browsers. Cytoscape.js has two ways to represent network data, and currently both reader and writer support only the array style graph notation. For example, this network in Cytoscape: {{attachment:JSON1.png}} will be exported to this JSON: {{{ { "elements" : { "nodes" : [ { "data" : { "id" : "723", "selected" : false, "annotation_Taxon" : "Saccharomyces cerevisiae", "alias" : [ "RPL31A", "RPL34", "S000002233", "ribosomal protein L31A (L34A) (YL28)" ], "shared_name" : "YDL075W", "SUID" : 723, "degree_layout" : 1, "name" : "YDL075W" }, "position" : { "x" : 693.0518315633137, "y" : -49.47506554921466 }, "selected" : false }, { "data" : { "id" : "726", "selected" : false, "annotation_Taxon" : "Saccharomyces cerevisiae", "alias" : [ "RP23", "RPL16B", "S000005013", "ribosomal protein L16B (L21B) (rp23) (YL15)" ], "shared_name" : "YNL069C", "SUID" : 726, "degree_layout" : 1, "name" : "YNL069C" }, "position" : { "x" : 627.3147710164387, "y" : -205.99251969655353 }, "selected" : false }, { "data" : { "id" : "658", "selected" : false, "annotation_Taxon" : "Saccharomyces cerevisiae", "alias" : [ "RPL11B", "S000003317", "ribosomal protein L11B (L16B) (rp39B) (YL22)" ], "shared_name" : "YGR085C", "SUID" : 658, "degree_layout" : 2, "name" : "YGR085C" }, "position" : { "x" : 804.3092778523762, "y" : -245.6235926946004 }, "selected" : false }, { "data" : { "id" : "660", "selected" : false, "annotation_Taxon" : "Saccharomyces cerevisiae", "alias" : [ "KAP108", "S000002803", "SXM1" ], "shared_name" : "YDR395W", "SUID" : 660, "degree_layout" : 8, "name" : "YDR395W" }, "position" : { "x" : 730.8733342488606, "y" : -157.50702317555744 }, "selected" : false }, { "data" : { "id" : "579", "selected" : false, "annotation_Taxon" : "Saccharomyces cerevisiae", "alias" : [ "RPL11A", "S000006306", "ribosomal protein L11A (L16A) (rp39A) (YL22)" ], "shared_name" : "YPR102C", "SUID" : 579, "degree_layout" : 2, "name" : "YPR102C" }, "position" : { "x" : 841.1395696004231, "y" : -130.77909119923908 }, "selected" : false }, { "data" : { "id" : "578", "selected" : false, "annotation_Taxon" : "Saccharomyces cerevisiae", "alias" : [ "GRC5", "QSR1", "RPL10", "S000004065", "ribosomal protein L10" ], "shared_name" : "YLR075W", "SUID" : 578, "degree_layout" : 2, "name" : "YLR075W" }, "position" : { "x" : 910.3755162556965, "y" : -217.0562556584676 }, "selected" : false } ], "edges" : [ { "data" : { "id" : "659", "source" : "658", "target" : "578", "selected" : false, "interaction" : "pp", "shared_interaction" : "pp", "shared_name" : "YGR085C (pp) YLR075W", "SUID" : 659, "name" : "YGR085C (pp) YLR075W" }, "selected" : false }, { "data" : { "id" : "661", "source" : "658", "target" : "660", "selected" : false, "interaction" : "pp", "shared_interaction" : "pp", "shared_name" : "YGR085C (pp) YDR395W", "SUID" : 661, "name" : "YGR085C (pp) YDR395W" }, "selected" : false }, { "data" : { "id" : "724", "source" : "660", "target" : "723", "selected" : false, "interaction" : "pp", "shared_interaction" : "pp", "shared_name" : "YDR395W (pp) YDL075W", "SUID" : 724, "name" : "YDR395W (pp) YDL075W" }, "selected" : false }, { "data" : { "id" : "733", "source" : "660", "target" : "579", "selected" : false, "interaction" : "pp", "shared_interaction" : "pp", "shared_name" : "YDR395W (pp) YPR102C", "SUID" : 733, "name" : "YDR395W (pp) YPR102C" }, "selected" : false }, { "data" : { "id" : "727", "source" : "660", "target" : "726", "selected" : false, "interaction" : "pp", "shared_interaction" : "pp", "shared_name" : "YDR395W (pp) YNL069C", "SUID" : 727, "name" : "YDR395W (pp) YNL069C" }, "selected" : false }, { "data" : { "id" : "580", "source" : "578", "target" : "579", "selected" : false, "interaction" : "pp", "shared_interaction" : "pp", "shared_name" : "YLR075W (pp) YPR102C", "SUID" : 580, "name" : "YLR075W (pp) YPR102C" }, "selected" : false } ] } } }}} And this is a sample visualization in Cytoscape.js: {{attachment:JSON2.png}} ==== Important Note ==== Export network and table to Cytoscape.js feature in Cytoscape creates a JSON file '''WITHOUT''' style. This means that you need to export the style in a separate JSON file if you apply style to your network. Please read Style section for more details. |
#DEPRECATED <<Include(Cytoscape_3/UserManual/Deprecated)>> |
This is a legacy document
This page has been deprecated and is no longer updated. The current version of the Cytoscape manual can be found at http://manual.cytoscape.org/
- Simple interaction file (SIF or .sif format)
- Nested network format (NNF or .nnf format)
- Graph Markup Language (GML or .gml format)
- XGMML (extensible graph markup and modelling language).
- SBML
- BioPAX
- PSI-MI Level 1 and 2.5
- GraphML
- Delimited text
- Excel Workbook (.xls, .xlsx)
The SIF format specifies nodes and interactions only, while other formats store additional information about network layout and allow network data exchange with a variety of other network programs and data sources. Typically, SIF files are used to import interactions when building a network for the first time, since they are easy to create in a text editor or spreadsheet. Once the interactions have been loaded and network layout has been performed, the network may be saved to GML or XGMML format for interaction with other systems. All file types listed (except Excel) are text files and you can edit and view them in a regular text editor.
SIF Format
The simple interaction format is convenient for building a graph from a list of interactions. It also makes it easy to combine different interaction sets into a larger network, or add new interactions to an existing data set. The main disadvantage is that this format does not include any layout information, forcing Cytoscape to re-compute a new layout of the network each time it is loaded.
Lines in the SIF file specify a source node, a relationship type (or edge type), and one or more target nodes:
nodeA <relationship type> nodeB nodeC <relationship type> nodeA nodeD <relationship type> nodeE nodeF nodeB nodeG ... nodeY <relationship type> nodeZ
A more specific example is:
node1 typeA node2 node2 typeB node3 node4 node5 node0
The first line identifies two nodes, called node1 and node2, and a single relationship between node1 and node2 of type typeA. The second line specifies three new nodes, node3, node4, and node5; here "node2" refers to the same node as in the first line. The second line also specifies three relationships, all of type typeB and with node2 as the source, with node3, node4, and node5 as the targets. This second form is simply shorthand for specifying multiple relationships of the same type with the same source node. The third line indicates how to specify a node that has no relationships with other nodes. This form is not needed for nodes that do have relationships, since the specification of the relationship implicitly identifies the nodes as well.
Duplicate entries are ignored. Multiple edges between the same nodes must have different edge types. For example, the following specifies two edges between the same pair of nodes, one of type xx and one of type yy:
node1 xx node2 node1 xx node2 node1 yy node2
Edges connecting a node to itself (self-edges) are also allowed:
node1 xx node1
Every node and edge in Cytoscape has a name column. For a network defined in SIF format, node names should be unique, as identically named nodes will be treated as identical nodes. The name of each node will be the name in this file by default (unless another string is mapped to display on the node using styles). This is discussed in the section on Styles. The name of each edge will be formed from the name of the source and target nodes plus the interaction type: for example, sourceName (edgeType) targetName.
The tag <edgeType> can be any string. Whole words or concatenated words may be used to define types of relationships, e.g. geneFusion, cogInference, pullsDown, activates, degrades, inactivates, inhibits, phosphorylates, upRegulates, etc.
Some common interaction types used in the Systems Biology community are as follows:
pp .................. protein – protein interaction pd .................. protein -> DNA (e.g. transcription factor binding upstream of a regulating gene.)
Some less common interaction types used are:
pr .................. protein -> reaction rc .................. reaction -> compound cr .................. compound -> reaction gl .................. genetic lethal relationship pm .................. protein-metabolite interaction mp .................. metabolite-protein interaction
Delimiters
Whitespace (space or tab) is used to delimit the names in the simple interaction file format. However, in some cases spaces are desired in a node name or edge type. The standard is that, if the file contains any tab characters, then tabs are used to delimit the fields and spaces are considered part of the name. If the file contains no tabs, then any spaces are delimiters that separate names (and names cannot contain spaces).
If your network unexpectedly contains no edges and node names that look like edge names, it probably means your file contains a stray tab that's fooling the parser. On the other hand, if your network has nodes whose names are half of a full name, then you probably meant to use tabs to separate node names with spaces.
Networks in simple interactions format are often stored in files with a .sif extension, and Cytoscape recognizes this extension when browsing a directory for files of this type.
NNF
The NNF format is a very simple format that unlike SIF allows the optional assignment of single nested network per node. No other node columns can be specified. There are only 2 possible line formats:
- A node "node" contained in a "network:"
network node
- 2 nodes linked together contained in a network:
network node1 interaction node2
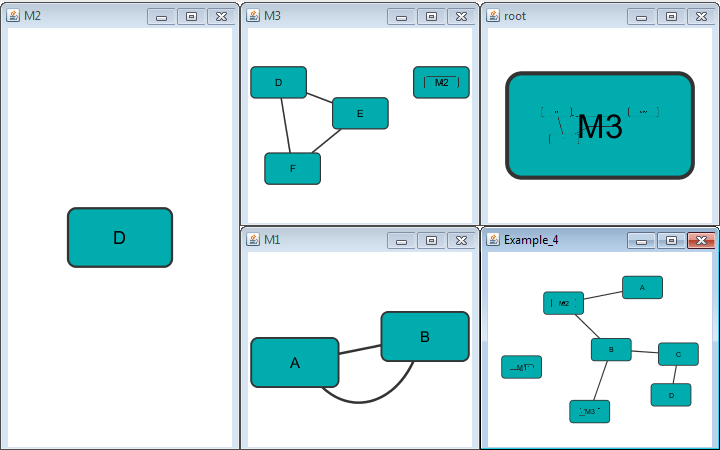
If a network name (first entry on a line) appeared previously as a node name (in columns 2 or 4), the network will be nested in the node with the same name. Also, if a name that has been previously defined as a network (by being listed in the first column), later appears as a node name (in columns 2 or 4), the previously defined network will be nested in the node with the same name. In summary: any time a name is used as both, a network name , and a node name, this implies that the network will be nested in the node of the same name. Additionally comments may be included on all lines. Comments start with a hash mark '#' and continue to the end of a line. Trailing comments (after data lines) and entirely blank lines anywhere are also permissible. Please note that if you load multiple NNF files in Cytoscape they will be treated like a single, long concatenated NNF file! If you need to embed spaces, tabs or backslashes in a name, you must escape it by preceding it with a backslash, so that, e.g. an embedded backslash becomes two backslashes, an embedded space a backslash followed by a space etc.
Examples
Example 1
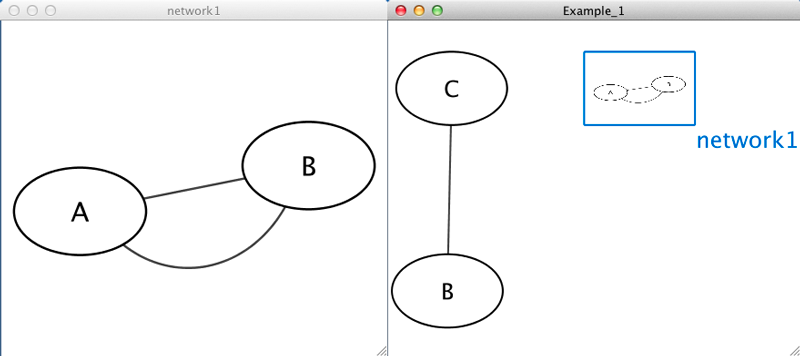
Example_1 C Example_1 network1 network1 A pp B network1 B pp A Example_1 C pp B
Example 2
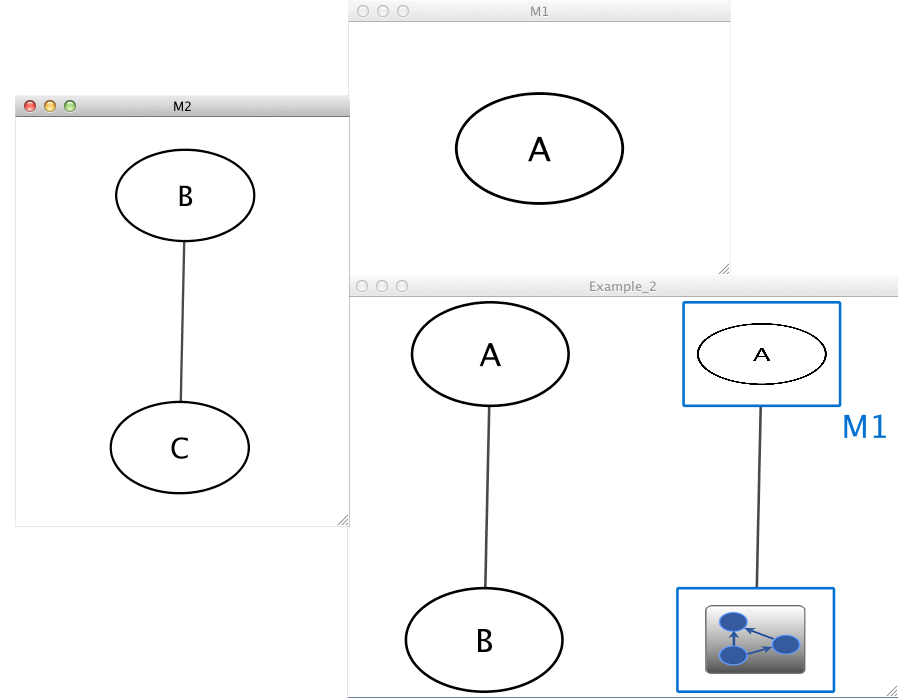
Example_2 M1 Example_2 M2 M1 A M2 B pp C Example_2 A pp B Example_2 M1 im M2
Example 3
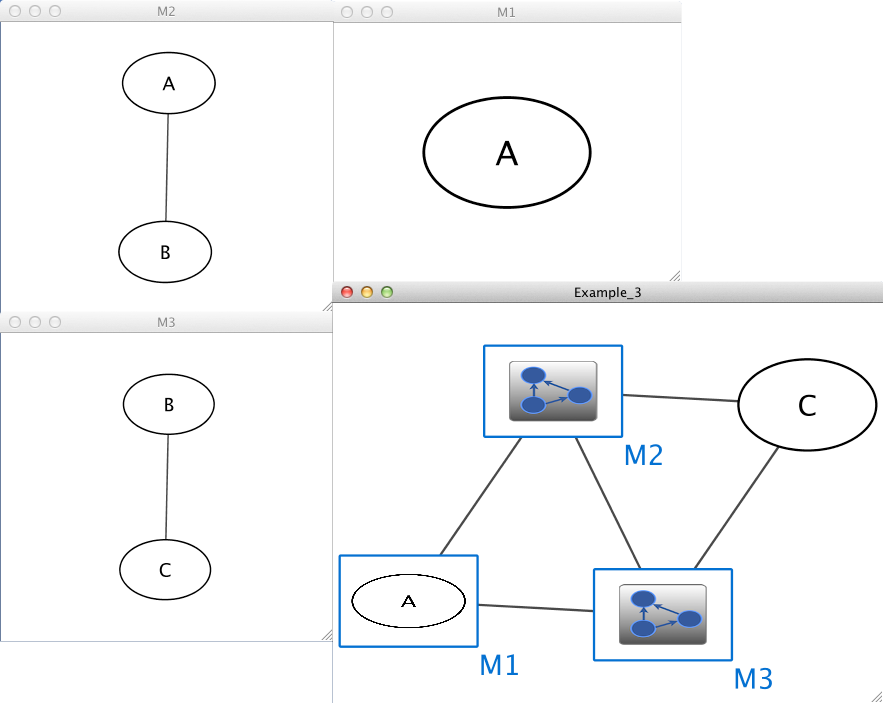
Example_3 M1 im M2 Example_3 M3 im M1 Example_3 M2 im M3 Example_3 C pp M3 Example_3 M2 pp C M1 A M2 A pp B M3 B pp C
Example 5
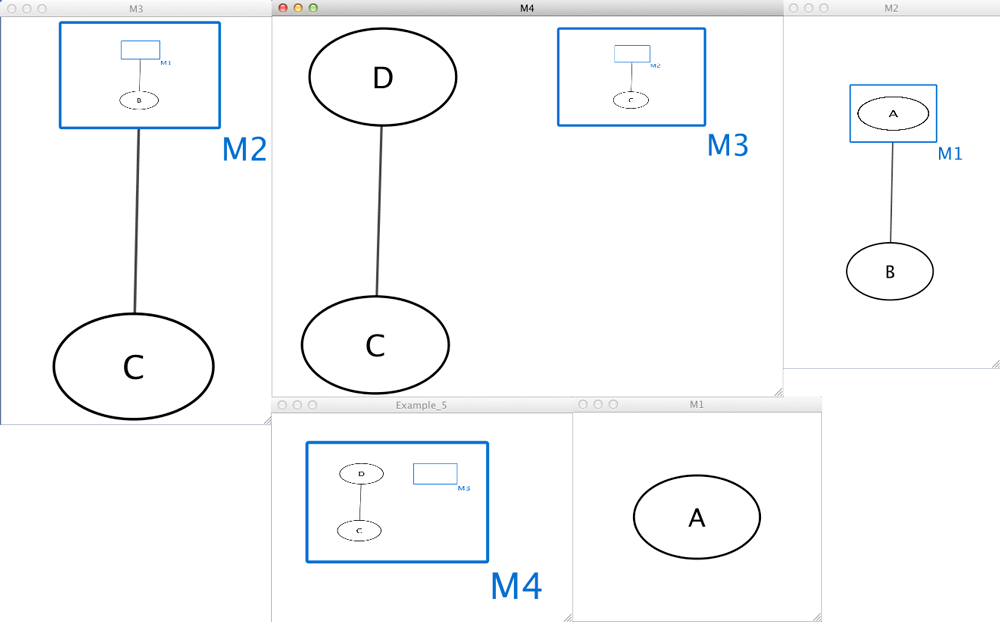
Example_5 M4 M4 D M4 M3 M3 M2 pp C M2 M1 pp B M1 A M4 C pp D
GML Format
In contrast to SIF, GML is a rich graph format language supported by many other network visualization packages. The GML file format specification is available at:
http://www.infosun.fmi.uni-passau.de/Graphlet/GML/
It is generally not necessary to modify the content of a GML file directly. Once a network is built in SIF format and then laid out, the layout is preserved by saving to and loading from GML. Properties specified in a GML file will result in a new style named Filename.style when that GML file is loaded.
XGMML Format
XGMML is the XML evolution of GML and is based on the GML definition. In addition to network data, XGMML contains node/edge/network column data. The XGMML file format specification is available at:
http://cgi5.cs.rpi.edu/research/groups/pb/punin/public_html/XGMML/
XGMML is now preferred to GML because it offers the flexibility associated with all XML document types. If you're unsure about which to use, choose XGMML.
There is a java system property "cytoscape.xgmml.repair.bare.ampersands" that can be set to "true" if you have experience trouble reading older files.
This should only be used when an XGMML file or session cannot be read due improperly encoded ampersands, as it slows down the reading process, but this is still preferable to attempting to fix such files using manual editing.
SBML (Systems Biology Markup Language) Format
The Systems Biology Markup Language (SBML) is an XML format to describe biochemical networks. SBML file format specification is available at:
BioPAX (Biological PAthways eXchange) Format
BioPAX is an OWL (Web Ontology Language) document designed to exchange biological pathways data. The complete set of documents for this format is available at:
PSI-MI Format
The PSI-MI format is a data exchange format for protein-protein interactions. It is an XML format used to describe PPI and associated data. PSI-MI XML format specification is available at:
http://psidev.sourceforge.net/mi/xml/doc/user/
GraphML
GraphML is a comprehensive and easy-to-use file format for graphs. It is based on XML. The complete set of documents for this format is available at:
http://graphml.graphdrawing.org/
Delimited Text Table and Excel Workbook
Cytoscape has native support for Microsoft Excel files (.xls, .xlsx) and delimited text files. The tables in these files can have network data and edge columns. Users can specify columns containg source nodes, target nodes, interaction types, and edge columns during file import. Some of the other network analysis tools, such as igraph (http://cneurocvs.rmki.kfki.hu/igraph/), has feature to export graph as simple text files. Cytoscape can read these text files and build networks from them. For more detail, please read the Import Free-Format Tables section of the Creating Networks section.
Cytoscape.js JSON
From Cytoscape 3.1.0 on, Cytoscape supports Cytoscape.js JSON files. You can use this feature to export your network visualizations to web browsers. Cytoscape.js has two ways to represent network data, and currently both reader and writer support only the array style graph notation. For example, this network in Cytoscape:
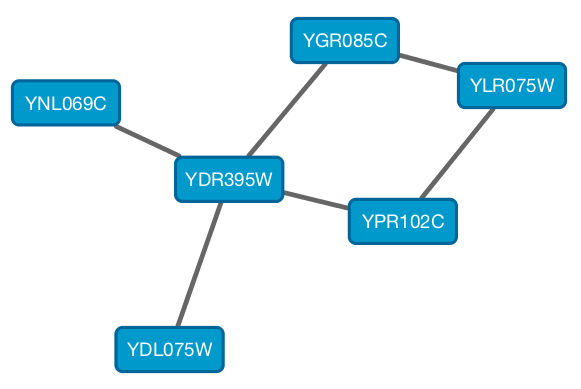
will be exported to this JSON:
{
"elements" : {
"nodes" : [ {
"data" : {
"id" : "723",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RPL31A", "RPL34", "S000002233", "ribosomal protein L31A (L34A) (YL28)" ],
"shared_name" : "YDL075W",
"SUID" : 723,
"degree_layout" : 1,
"name" : "YDL075W"
},
"position" : {
"x" : 693.0518315633137,
"y" : -49.47506554921466
},
"selected" : false
}, {
"data" : {
"id" : "726",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RP23", "RPL16B", "S000005013", "ribosomal protein L16B (L21B) (rp23) (YL15)" ],
"shared_name" : "YNL069C",
"SUID" : 726,
"degree_layout" : 1,
"name" : "YNL069C"
},
"position" : {
"x" : 627.3147710164387,
"y" : -205.99251969655353
},
"selected" : false
}, {
"data" : {
"id" : "658",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RPL11B", "S000003317", "ribosomal protein L11B (L16B) (rp39B) (YL22)" ],
"shared_name" : "YGR085C",
"SUID" : 658,
"degree_layout" : 2,
"name" : "YGR085C"
},
"position" : {
"x" : 804.3092778523762,
"y" : -245.6235926946004
},
"selected" : false
}, {
"data" : {
"id" : "660",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "KAP108", "S000002803", "SXM1" ],
"shared_name" : "YDR395W",
"SUID" : 660,
"degree_layout" : 8,
"name" : "YDR395W"
},
"position" : {
"x" : 730.8733342488606,
"y" : -157.50702317555744
},
"selected" : false
}, {
"data" : {
"id" : "579",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "RPL11A", "S000006306", "ribosomal protein L11A (L16A) (rp39A) (YL22)" ],
"shared_name" : "YPR102C",
"SUID" : 579,
"degree_layout" : 2,
"name" : "YPR102C"
},
"position" : {
"x" : 841.1395696004231,
"y" : -130.77909119923908
},
"selected" : false
}, {
"data" : {
"id" : "578",
"selected" : false,
"annotation_Taxon" : "Saccharomyces cerevisiae",
"alias" : [ "GRC5", "QSR1", "RPL10", "S000004065", "ribosomal protein L10" ],
"shared_name" : "YLR075W",
"SUID" : 578,
"degree_layout" : 2,
"name" : "YLR075W"
},
"position" : {
"x" : 910.3755162556965,
"y" : -217.0562556584676
},
"selected" : false
} ],
"edges" : [ {
"data" : {
"id" : "659",
"source" : "658",
"target" : "578",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YGR085C (pp) YLR075W",
"SUID" : 659,
"name" : "YGR085C (pp) YLR075W"
},
"selected" : false
}, {
"data" : {
"id" : "661",
"source" : "658",
"target" : "660",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YGR085C (pp) YDR395W",
"SUID" : 661,
"name" : "YGR085C (pp) YDR395W"
},
"selected" : false
}, {
"data" : {
"id" : "724",
"source" : "660",
"target" : "723",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YDR395W (pp) YDL075W",
"SUID" : 724,
"name" : "YDR395W (pp) YDL075W"
},
"selected" : false
}, {
"data" : {
"id" : "733",
"source" : "660",
"target" : "579",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YDR395W (pp) YPR102C",
"SUID" : 733,
"name" : "YDR395W (pp) YPR102C"
},
"selected" : false
}, {
"data" : {
"id" : "727",
"source" : "660",
"target" : "726",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YDR395W (pp) YNL069C",
"SUID" : 727,
"name" : "YDR395W (pp) YNL069C"
},
"selected" : false
}, {
"data" : {
"id" : "580",
"source" : "578",
"target" : "579",
"selected" : false,
"interaction" : "pp",
"shared_interaction" : "pp",
"shared_name" : "YLR075W (pp) YPR102C",
"SUID" : 580,
"name" : "YLR075W (pp) YPR102C"
},
"selected" : false
} ]
}
}And this is a sample visualization in Cytoscape.js:
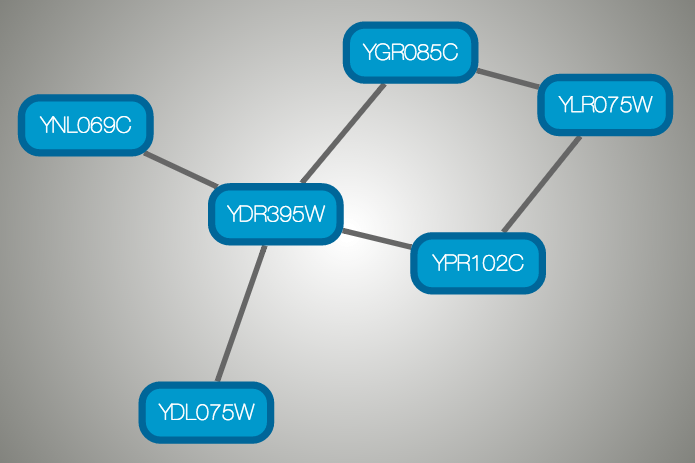
Important Note
Export network and table to Cytoscape.js feature in Cytoscape creates a JSON file WITHOUT style. This means that you need to export the style in a separate JSON file if you apply style to your network. Please read Style section for more details.